the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
DNA methylation pattern of the goat PITX1 gene and its effects on milk performance
Haiyu Zhao
Sihuan Zhang
Xianfeng Wu
Chuanying Pan
Xiangchen Li
Chuzhao Lei
Hong Chen
Xianyong Lan
Paired-like homeodomain transcription factor 1 (PITX1) is a pivotal gene in the hypothalamic–pituitary–adrenal axis, which is a well-known pathway affecting lactation performance. The aim of this study was to analyze the DNA methylation profile of the PITX1 gene and its relevance to milk performance in Xinong Saanen dairy goats; thus, potential epigenetic markers of lactation performance were identified. A total of 267 goat blood samples were divided into “low” and “high” groups according to two milk traits: the average milk yield (AMY) and the average milk density (AMD). One CpG island in the 3′-flanking region of the PITX1 gene was identified as being related to milk performance. Fisher's exact test demonstrated that the methylation rates of the overall CpG island and the 3rd and 12th CpG-dinucleotide loci in the blood were significantly associated with the AMY, and the overall methylation rate of the high AMY group was relative hypomethylation compared with the low AMY group. The overall methylation rates of this CpG island in mammary gland tissue from dry and lactation periods again exhibited a significant difference: the lactation period showed relative hypomethylation compared with the dry period. Bioinformatic transcription factor binding site prediction identified some lactation performance related transcription factors in this CpG island, such as CTCF, STAT, SMAD, CDEF, SP1, and KLFS. Briefly, overall methylation changes of the CpG island in the PITX1 gene are relevant to lactation performance, which will be valuable for future studies and epigenetic marker-assisted selection (eMAS) in the breeding of goats with respect to lactation performance.
- Article
(466 KB) - Full-text XML
- BibTeX
- EndNote
Lactation performance is a complex quantitative trait affected by several crucial factors including genetic background, nutrition, and environment (Carcangiu et al., 2018). Over the past decade, the influences of genetic variations on milk performance in animals have been extensively studied (Koufariotis et al., 2018). Recently an increasing number of studies have demonstrated that nutritional or environmental changes contribute to changes in inheritable epigenetic modifications, such as DNA methylation (Lan et al., 2013a; Chavatte-Palmer et al., 2018; Kwan et al., 2018; Yuan et al., 2018). As one of the most important and common types of epigenetic modification, DNA methylation plays a significant role in regulating biological processes such as milk production and organ development (Hwang et al., 2017; Fleming et al., 2018; Jessop et al., 2018). In recent years, gene methylation involved in vital biological functions has been well-studied in animals (Ibeagha-Awemu and Zhao, 2015). However, to date, no association has been established between epigenetic modifications and lactation performance.
It is well-known that lactation performance is regulated by several critical pathways, such as the hypothalamic–pituitary–adrenal (HPA) axis pathway (PITX2/PITX1–HESX1–LHX3/LHX4–PROP1–POU1F1), which regulates the expression of GH (growth hormone) and PRL (prolactin) (Davis et al., 2010). As a member of the paired-like homeodomain transcription factor (PITX) family, the PITX1 gene is located upstream of the HPA axis pathway and plays a critical role in pituitary organogenesis (Ma et al., 2017). During early pituitary development, PITX1 activates the transcription of couples of pituitary hormone genes and transcription factor genes such as the LIM family, the POU family, and the SIX family (Poulin et al., 2000). Notably, PITX1 has been characterized as an activator of the POMC gene and a partner of Pit1 causing differential expression of GH, PRL, and TSH, and thereby affecting milk performance in animals (Carvalho et al., 2006). In 2013, both the PITX1 gene and another PITX family member, PITX2, were identified as being associated with milk performance in dairy goats (Lan et al., 2013b; Zhao et al., 2013).
Therefore, this study aimed at exploring the DNA methylation changes at the CpG islands within PITX1 gene, as well as their relevance to milk performance. The present study is important and necessary for exploring the epigenetic regulation of lactation and developing potential epigenetic markers for improving milk performance through epigenetic marker-assisted selection (eMAS) in breeding.
Experimental animals and the procedures performed in this study were approved by the Faculty Animal Policy and Welfare Committee of Northwest A & F University under contract (NWAFU-314020038). The care and use of experimental animals fully complied with local animal welfare laws, guidelines, and policies.
2.1 Animals, phenotypic data recording, and group classification
A total of 267 blood samples were obtained from healthy, unrelated adult female Xinong Saanen dairy goats, which were reared on the Chinese native dairy goat breeding farm in Qianyang County, Shaanxi, China (Zhao et al., 2013). The data regarding the average milk yield (AMY; kg) were obtained directly from the breeding farm, whereas the data regarding the average milk density (AMD) were measured by our study team using a MilkoScan FT120 (FOSS Corporation, Denmark) instrument (Lan et al., 2013b; Zhao et al., 2013). The relevant measurements of these two milk traits are listed in Table 1.
Table 1Milk performance of the low and high groups of Xinong Saanen dairy goats.
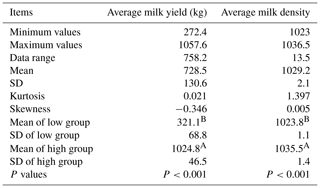
The use of different letters (A, B) beside mean values in the same column signifies a significant difference at the P<0.001 level; SD stands for standard deviation.
Using the statistical probability (95.00 %) of the mean ±1.96 standard deviations for AMY and AMD, the low and high groups for each of the aforementioned milk traits were classified according to published literature (Pan et al., 2013). For the low or high group of each milk trait, two random pooled samples were constructed, and were named as follows: L-P1 (low-pool I), L-P2 (low-pool II), H-P1 (high-pool I), and H-P2 (high-pool II). Finally, eight DNA pools were constructed. It can be noted from Table 1 that the low group for each milk trait was significantly lower than average, whereas the high group was significantly higher than average. A t test was used to confirm the significance of the difference between the low and high groups, and the results validated that there were indeed significant differences between low and high groups for the AMY (kg) and the AMD (P<0.001; Table 1). In addition, five mammary gland tissue samples from Xinong Saanen dairy goats during a lactation period (n=2) and a dry period (n=3) were collected from another Xinong Saanen dairy goat farm, Fuping, Shaanxi Province, China (X. Zhang et al., 2017).
2.2 Genomic DNA extraction and bisulfite treatment
Genomic DNA samples from all chosen individuals were isolated as previously described (Cui et al., 2018; Wang et al., 2018). The DNA samples were then measured and diluted to a final concentration of 50 ng µL−1 (Wang et al., 2017; Q. Yang et al., 2017, 2018). Qualified DNA samples from the low and high groups were equally selected to construct genomic DNA pools. Each pool – containing 1000 ng mixed genomic DNA – was processed by sodium bisulfite using a QIAGEN EpiTect Fast DNA Bisulfite Kit (QIAGEN, Germany) and following the manufacture's instructions.
2.3 Bioinformatics analyses
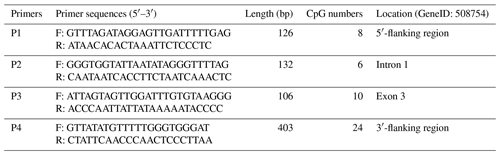
The CpG islands within the goat PITX1 gene were predicted using the MethPrimer website (http://www.urogene.org/methprimer/, last access: 10 January 2019) (Li and Dahiya, 2002), and the primers used to amplify the CpG islands were also designed using this website (Table 2).
To identify the possible trans-acting factors which might bind to specific CpG-dinucleotide locus, two respective authorized and reliable online bioinformatics softwares were used: TFSEARCH (Version 1.3) (http://mbs.cbrc.jp/research/db/TFSEARCH.html, last access: 19 June 2018) and the MatInspector database in Genomatix (http://www.genomatix.de, last access: 10 January 2019).
2.4 PCR amplification, cloning, and sequencing of the CpG island within the PITX1 gene
The PCR reaction and the purification of the PCR products were performed as previously described (Pan et al., 2013). Purified fragments were cloned into the pGEM-T Easy Vector (Promega, WI, USA) system. The colony PCR was used to identify the positive inserts, which were sequenced via sequencing service (GenScript, Nanjing, China). The number of positive inserts for each pool sent for sequencing was around 10, according to the efficiency of ligation and transformation. The methylation status for the CpG island and each CpG-dinucleotide locus was measured by sequencing, and alignment analyses were carried out using DNAMAN software (version 7.0, Lynnon Biosoft, Vaudreuil, Quebec, Canada) (Lan et al., 2013a).
2.5 Statistical analysis
Comparisons of the methylation difference at each CpG-dinucleotide locus, the entire CpG island between the low and high groups for each milk trait, and the methylation status of mammary gland tissue from dry and lactation periods were analyzed using the Fisher's exact test (χ2-test) as previously described (Lan et al., 2013a).
3.1 DNA methylation profile of the PITX1 gene
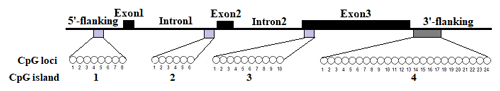
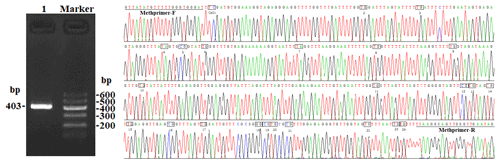
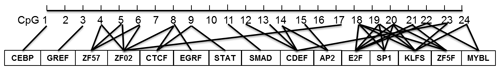
In this study, four CpG islands located at the 5′-flanking region, intron 1, exon 3, and 3′-flanking region of the PITX1 gene were predicted (Fig. 1). A total of four pairs of methylation primers were designed to amplify these four CpG islands. Due to the complexity and low G and C bases content of bisulfite-processed genomic DNA as well as the low amplification efficiency for bisulfite PCR, only one 403 bp-CpG island (24 CpG-dinucleotide loci) located at the 3′-flanking region was accessible using the P4 primer (Fig. 2). At the available CpG island within the PITX1 gene, about 10 positive inserts for each sample were sequenced. The alignments of the sequences demonstrated different overall methylation patterns in the low and high pools for the two milk traits; the overall methylation rate of these blood DNA pools varied from 22.46 % (high pool group for AMY) to 33.33 % (high pool group for the AMD) (Table 3). In addition, different overall methylation patterns were detected in mammary gland tissue from dry (57.29 %) and lactation (45.03 %) periods (Table 4).
Table 3Methylation percentage and P value difference of each CpG-dinucleotide locus between the low and high groups.
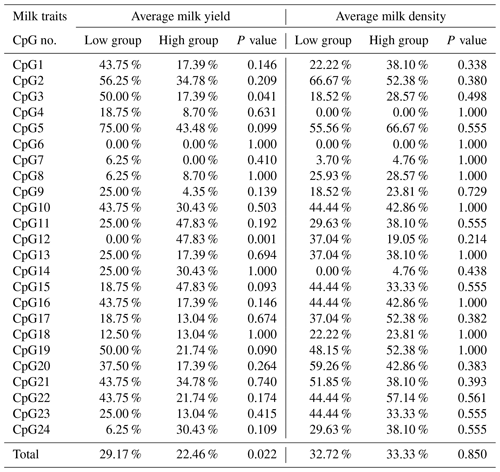
3.2 DNA methylation differences between the low and high groups for the two milk traits
Based on the methylation status of each CpG-dinucleotide locus, methylation differences between the low and high groups were evaluated according to Fisher's exact test (χ2-test). As seen from Table 3 and Fig. 3, the overall methylation level of the CpG island within the PITX1 gene was significantly associated with the AMY (P=0.022). Specifically, the overall methylated percentage of the high AMY group (22.46 %) was significantly lower than that of the low group (29.17 %). For individual CpG-dinucleotide locus, the methylation status of the 3rd CpG-dinucleotide locus was significantly associated with the AMY (P=0.041), and the methylation percentage of this locus from high group (17.39 %) was significantly lower than that from low group (50.00 %); on the contrary, the methylation status of the 12th CpG-dinucleotide locus showed a negative association with the AMY, and the methylation rate of the high group (47.83 %) was significantly higher than that of the low group (0 % (Table 3 and Fig. 3). However, no significant relevance was found between methylation changes and the AMD.
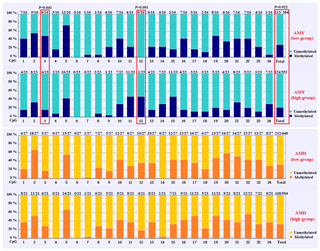
Figure 3Comparisons of the DNA methylation differences for each CpG locus between the low and high groups for the two lactation traits. The numbers (X∕Y) above each bar represent the ratio of methylation versus total colonies, and the red frames represent significant differences. AMY refers to the average milk yield, and AMD refers to the average milk density.
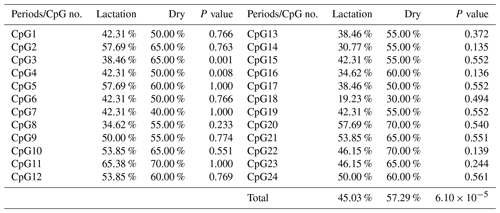
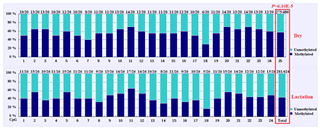
3.3 DNA methylation differences between mammary gland tissue from dry and lactation periods
The overall DNA methylation rates of the CpG island in mammary gland tissue from dry and lactation periods exhibited a significant difference (); mammary gland tissue from the dry period showed hypermethylation (57.29 %), whereas mammary gland tissue from the lactation period showed relative hypomethylation (45.03 %) (Table 4 and Fig. 4). However, no significant relevance was identified on each CpG-dinucleotide locus in mammary gland tissue from the dry and lactation periods.
3.4 Possible trans-acting factors binding to the CpG-dinucleotide loci
According to the bioinformatical analyses, several crucial transcription factors, such as CTCF, STAT, SMAD, CDEF, SP1, KLFS, and zinc finger transcription factors were predicted to bind to the CpG-dinucleotide loci of this CpG island within PITX1 (Fig. 5). In this CpG island, 24 CpG-dinucleotide loci were sparsely localized in the 403 bp region, and potential transcription factors were predicted to bind to the nearby area of almost every CpG-dinucleotide loci.
Extensive DNA methylation studies have focused on the CpG islands located at the promoter region, as the methylation of promoter CpG islands could result in self-perpetuating gene silencing (Moore et al., 2013). However, an increasing number of studies have found that a far greater proportion of methylation occurs across the whole gene (Jones, 2012). Genome-wide studies have revealed that gene body methylation is evolutionarily conserved (Keller et al., 2016). Therefore, this work attempted to study the methylation of four possible CpG islands across the whole PITX1 gene in a group Xinong Saanen dairy goats. However, only the CpG island located at the 3′-flanking region was available. It is well known that 3′-flanking regions are also important in gene regulation (Yang et al., 2016).
Recently, numerous epigenetic markers have been identified as being involved in biological functions and livestock traits using whole genome bisulfite sequencing (WGBS) and bisulfite sequencing methods (Y. Yang et al., 2017; Sarova et al., 2018). Fang et al. (2017) uncovered 331 meat quality traits related to candidate differentially methylated regions (DMRs) in Japanese Black (Wagyu) and Chinese Red Steppes cattle. Y. Zhang et al. (2017) revealed 10 DMR-related genes (DMGs) by analyzing the sequencing results of Hu sheep ovaries, which were likely related to the prolificacy of Hu sheep. Li et al. (2018) identified potential DMRs and DMGs that were involved in hair follicle development and growth in cashmere goats. Dechow and Liu (2018) detected the DNA methylation patterns in peripheral blood mononuclear cells of Holstein cattle with variable milk yield, and revealed some potential epigenetic variation related to milk performance. With the rapid development of science, more and more crucial genetic and epigenetic markers will be discovered, which will provide abundant information for animal breeding and biological research.
Numerous genes have been proved to be regulated by DNA methylation, and there are a few possible mechanisms by which this can occur (Mattern et al., 2017). The methyl-group in CpG islands might result in compact and inactive chromatin structure, recruiting methyl-CpG binding proteins or preventing relevant transcription factors from binding, which inhibits the initiation or proceeding of transcription (Kang et al., 2015). In this study, we designed three pairs of primers to detect the expression of goat PITX1. However, we failed to detect the expression of goat PITX1 in mammary gland tissue and other tissues (heart, liver, spleen, lung, kidney, muscle, and adipose tissue – data not shown). Furthermore, according to the Ensembl, NCBI, and BioGPS databases, there is little expression of PITX1 in the adult mammary glands of humans, mice, and sheep. Even though there was no expression of PITX1 in mammary gland tissue, we speculate that the epigenetic modification of PITX1 can still be an useful epigenetic marker for improving lactation performance in the process of goat epigenetic marker-assisted selection in breeding at the DNA level.
In addition, these findings have also been corroborated by the biological functions of some possible transcription factors predicted in this study, including the following: (1) CTCF can regulate gene expression by binding to the gene imprinting control region (ICR) (Dunn and Davie, 2003); (2) the STAT family of proteins are the main components in the JAK–STAT pathway, which is critical to the proper development and function of the mammary epithelial tissue (Haricharan and Li, 2014); (3) SMAD proteins are Smad signaling components – in mammary epithelial cells, Smad signaling acts to antagonize JAK–STAT signaling to regulate mammary gland differentiation (Cocolakis et al., 2008); (4) CDEF could regulate the cell cycle of epithelial cell (Müller and Engeland, 2010); and (5) SP1 and KLFS could regulate the expression of genes containing GC-rich promoters (Kaczynski et al., 2003), and these transcription factors could influence the function of PITX1 and therefore affect lactation performance.
In this study, a functional CpG island at the 3′ flanking region of goat PITX1 gene was identified. In this region, the overall methylation rate of the high AMY group showed relative hypomethylation compared with the low AMY yield group; the overall methylation rate of mammary gland tissue from the lactation period showed hypomethylation compared with the dry period. These results will provide epigenetic markers for improving lactation performance with regards to the process of goat epigenetic marker-assisted selection in breeding.
The original data are available upon request to the corresponding author.
| PITX | Paired-like homeodomain transcription factor |
| PITX1 | Paired-like homeodomain transcription factor 1 |
| AR | Androgen receptor |
| eMAS | Epigenetic marker-assisted selection |
| HPA axis | Hypothalamic–pituitary–adrenal axis |
| SD | Standard deviation |
| AMY | Average milk yield |
| AMD | Average milk density |
| WGBS | Whole genome bisulfite sequencing |
| DMRs | Differentially methylated regions |
| DMGs | Differentially methylated region-related genes |
HZ and SZ contributed equally to this work.
The authors declare that they have no conflict of interest.
This work was supported by the Natural Science Foundation of China
(grant no. 31172184) and the Natural Science Foundation of Shaanxi Province of
China (grant no. 2017JM3021).
Edited by: Steffen
Maak
Reviewed by: Runjun Yang and one anonymous referee
Carcangiu, V., Arfuso, F., Luridiana, S., Giannetto, C., Rizzo, M., Bini, P. P., and Piccione, G.: Relationship between different livestock managements and stress response in dairy ewes, Arch. Anim. Breed., 61, 37–41, https://doi.org/10.5194/aab-61-37-2018, 2018.
Carvalho, L., Ward, R. D., Brinkmeier, M. L., Potok, M. A., Vesper, A. H., and Camper, S. A.: Molecular basis for pituitary dysfunction: comparison of Prop1 and Pit1 mutant mice, Dev. Biol., 295, 340, https://doi.org/10.1016/j.ydbio.2006.04.056, 2006.
Chavatte-Palmer, P., Velazquez, M. A., Jammes, H., and Duranthon, V.: Review: Epigenetics, developmental programming and nutrition in herbivores, Animal, 24, 1–9, 2018.
Cocolakis, E., Dai, M., Drevet, L., Ho, J., Haines, E., Ali, S., and Lebrun, J. J.: Smad signaling antagonizes STAT5-mediated gene transcription and mammary epithelial cell differentiation, J. Biol. Chem., 283, 1293–1307, 2008.
Cui, Y., Yan, H., Wang, K., Xu, H., Zhang, X., Zhu, H., Liu, J., Qu, L., Lan, X., and Pan, C.: Insertion/Deletion within the KDM6A gene is significantly associated with litter size in goat, Frontiers in Genetics, 9, 91, https://doi.org/10.3389/fgene.2018.00091, 2018.
Davis, S. W., Castinetti, F., Carvalho, L. R., Ellsworth, B. S., Potok, M. A., Lyons, R. H., Brinkmeier, M. L., Raetzman, L. T., Carninci, P., Mortensen, A. H., Hayashizaki, Y., Arnhold, I. J., Mendonça, B. B., Brue, T., and Camper, S. A.: Molecular mechanisms of pituitary organogenesis: In search of novel regulatory genes, Mol. Cell. Endocrinol., 323, 4–19, 2010.
Dechow, C. D. and Liu, W. S.: DNA methylation patterns in peripheral blood mononuclear cells from Holstein cattle with variable milk yield, BMC Genomics, 19, 744, https://doi.org/10.1186/s12864-018-5124-9, 2018.
Dunn, K. L. and Davie, J. R.: The many roles of the transcriptional regulator CTCF, Biochem. Cell Biol., 81, 161–167, 2003.
Fang, X., Zhao, Z., Yu, H., Li, G., Jiang, P., Yang, Y., Yang, R., and Yu, X.: Comparative genome-wide methylation analysis of longissimus dorsi muscles between Japanese black (Wagyu) and Chinese Red Steppes cattle, PLoS One, 12, e0182492, https://doi.org/10.1371/journal.pone.0182492, 2017.
Fleming, A., Abdalla, E. A., Maltecca, C., and Baes, C. F.: Invited review: Reproductive and genomic technologies to optimize breeding strategies for genetic progress in dairy cattle, Arch. Anim. Breed., 61, 43–57, https://doi.org/10.5194/aab-61-43-2018, 2018.
Haricharan, S. and Li, Y.: STAT signaling in mammary gland differentiation, cell survival and tumorigenesis, Mol. Cell. Endocrinol., 382, 560–569, 2014.
Hwang, J. H., An, S. M., Kwon, S., Park, D. H., Kim, T. W., Kang, D. G., Yu, G. E., Kim, I. S., Park, H. C., Ha, J., and Kim, C. W.: DNA methylation patterns and gene expression associated with litter size in Berkshire pig placenta, PLoS One, 12, e0184539, https://doi.org/10.1371/journal.pone.0184539, 2017.
Ibeagha-Awemu, E. M. and Zhao, X.: Epigenetic marks: regulators of livestock phenotypes and conceivable sources of missing variation in livestock improvement programs, Frontiers in Genetics, 6, 302, https://doi.org/10.3389/fgene.2015.00302, 2015.
Jessop, P., Ruzov, A., and Gering, M.: Developmental Functions of the Dynamic DNA Methylome and Hydroxymethylome in the Mouse and Zebrafish: Similarities and Differences, Frontiers in Cell and Developmental Biology, 6, 27, https://doi.org/10.3389/fcell.2018.00027, 2018.
Jones, P. A.: Functions of DNA methylation: islands, start sites, gene bodies and beyond, Nat. Rev. Genet., 13, 484–492, 2012.
Kaczynski, J., Cook, T., and Urrutia, R.: Sp1-and Kruppel-like transcription factors, Genome Biol., 4, 206, https://doi.org/10.1186/gb-2003-4-2-206, 2003.
Kang, J. Y., Song, S. H., Yun, J., Jeon, M. S., Kim, H. P., Han, S. W., and Kim, T. Y.: Disruption of CTCF/cohesin-mediated high-order chromatin structures by DNA methylation downregulates PTGS2 expression, Oncogene, 34, 5677–5684, 2015.
Keller, T. E., Han, P., and Yi, S. V.: Evolutionary Transition of Promoter and Gene Body DNA Methylation across Invertebrate-Vertebrate Boundary, Mol. Biol. Evol., 33, 1019–1028, 2016.
Koufariotis, L. T., Chen, Y. P., Stothard, P., and Hayes, B. J.: Variance explained by whole genome sequence variants in coding and regulatory genome annotations for six dairy traits, BMC Genomics, 19, 237, https://doi.org/10.1186/s12864-018-4617-x, 2018.
Kwan, S. T. C., King, J. H., Grenier, J. K., Yan, J., Jiang, X., Roberson, M. S., and Caudill, M. A.: Maternal Choline Supplementation during Normal Murine Pregnancy Alters the Placental Epigenome: Results of an Exploratory Study, Nutrients, 10, E417, https://doi.org/10.3390/nu10040417, 2018.
Lan, X. Y., Cretney, E. C., Kropp, J., Khateeb, K., Berg, M. A., Peñagaricano, F., and Khatib, H.: Maternal diet during pregnancy induces gene expression and DNA methylation changes in fetal tissues in sheep, Frontiers in Genetics, 4, 49, https://doi.org/10.3389/fgene.2013.00049, 2013a.
Lan, X. Y., Zhao, H. Y., Li, Z. J., Zhou, R., Pan, C. Y., Lei, C. Z., and Chen, H.: Exploring the Novel Genetic Variant of PITX1 Gene and Its Effect on Milk Performance in Dairy Goats, J. Integr. Agr., 2, 118–126, 2013b.
Li, C., Li, Y., Zhou, G., Gao, Y., Ma, S., Chen, Y., Song, J., and Wang, X.: Whole-genome bisulfite sequencing of goat skins identifies signatures associated with hair cycling, BMC Genomics, 19, 638, https://doi.org/10.1186/s12864-018-5002-5, 2018.
Li, L. C. and Dahiya, R.: MethPrimer: designing primers for methylation PCRs, Bioinformatics, 18, 1427–143, 2002.
Ma, L., Qin, Q. M., Yang, Q., Zhang, M., Zhao, H. Y., Pan, C. Y., Lei, C. Z., Chen, H., and Lan, X. Y.: Associations of six SNPs of POU1F1-PROP1-PITX1-SIX3 pathway genes with growth traits in two Chinese indigenous goat breeds, Ann. Anim. Sci., 17, 399–411, 2017.
Mattern, F., Heinzmann, J., Herrmann, D., Lucas-Hahn, A., Haaf, T., and Niemann, H.: Gene-specific profiling of DNA methylation and mRNA expression in bovine oocytes derived from follicles of different size categories, Reprod. Fert. Develop., 29, 2040–2051, 2017.
Moore, L. D., Le, T., and Fan, G.: DNA methylation and its basic function, Neuropsychopharmacology, 38, 23–38, 2013.
Müller, G. A. and Engeland, K.: The central role of CDE/CHR promoter elements in the regulation of cell cycle-dependent gene transcription, FEBS J., 277, 877–893, 2010.
Pan, C., Jia, W., Wu, X., Zhao, H., Liu, S., Lei, C., Lan, X., and Chen, H.: DNA methylation profile of DNA methyltransferase 3b (DNMT3b) gene and its influence on growth traits in goat, J. Anim. Plant Sci., 23, 380–387, 2013.
Poulin, G., Lebel, M., Chamberland, M., Paradis, F. W., and Drouin, J.: Specific protein-protein interaction between basic helix-loop-helix transcription factors and homeoproteins of the Pitx family, Mol. Cell. Biol., 20, 4826–4837, 2000.
Sarova, N., Ahlawat, S., Grewal, A., Sharma, R., and Arora, R.: Differential promoter methylation of DAZL gene in bulls with varying seminal parameters, Reprod. Domest. Anim., 53, 914–920, 2018.
Wang, X., Yang, Q., Wang, K., Zhang, S., Pan, C., Chen, H., Qu, L., Yan, H., and Lan, X.: A novel 12-bp indel polymorphism within the GDF9 gene is significantly associated with litter size and growth traits in goats, Anim. Genet., 48, 735–736, 2017.
Wang, X., Yang, Q., Wang, K., Yan, H., Pan, C., Chen, H., Liu, J., Zhu, H., Qu, L., and Lan, X.: Two strongly linked single nucleotide polymorphisms (Q320P and V397I) in GDF9 gene are associated with litter size in cashmere goats, Theriogenology, 125, 115–121, 2018.
Yang, Q., Zhang, S., Liu, L., Cao, X., Lei, C., Qi, X., Lin, F., Qu, W., Qi, X., Liu, J., Wang, R., Chen, H., and Lan, X.: Application of mathematical expectation (ME) strategy for detecting low frequency mutations: An example for evaluating 14-bp insertion/deletion (indel) within the bovine PRNP gene, Prion, 10, 409–419, 2016.
Yang, Q., Yan, H., Li, J., Xu, H., Wang, K., Zhu, H., Chen, H., Qu, L., and Lan, X.: A novel 14-bp duplicated deletion within goat GHR gene is significantly associated with growth traits and litter size, Anim. Genet., 48, 499–500, 2017.
Yang, Q., Zhang, S. L., Li, J., Wang, X., Peng, K., Lan, X., and Pan, C.: Development of a touch-down multiplex PCR method for simultaneously rapidly detecting three novel insertion/deletions (indels) within one gene: an example for goat GHR gene, Anim. Biotechnol., 1–6, https://doi.org/10.1080/10495398.2018.1517770, 2018.
Yang, Y., Liang, G., Niu, G., Zhang, Y., Zhou, R., Wang, Y., Mu, Y., Tang, Z., and Li, K.: Comparative analysis of DNA methylome and transcriptome of skeletal muscle in lean-, obese-, and mini-type pigs, Sci. Rep.-UK, 7, 39883, https://doi.org/10.1038/srep39883, 2017.
Yuan, X., Tsujimoto, K., Hashimoto, K., Kawahori, K., Hanzawa, N., Hamaguchi, M., Seki, T., Nawa, M., Ehara, T., Kitamura, Y., Hatada, I., Konishi, M., Itoh, N., Nakagawa, Y., Shimano, H., Takai-Igarashi, T., Kamei, Y., and Ogawa, Y.: Epigenetic modulation of Fgf21 in the perinatal mouse liver ameliorates diet-induced obesity in adulthood, Nat. Commun., 9, 636, https://doi.org/10.1038/s41467-018-03038-w, 2018.
Zhao, H., Wu, X., Cai, H., Pan, C., Lei, C., Chen, H., and Lan, X.: Genetic variants and effects on milk traits of the caprine paired-like homeodomain transcription factor 2 (PITX2) gene in dairy goats, Gene, 532, 203–210, 2013.
Zhang, X., Zhang, S., Ma, L., Jiang, E., Xu, H., Chen, R., Yang, Q., Chen, H., Li, Z., and Lan, X.: Reduced representation bisulfite sequencing (RRBS) of dairy goat mammary glands reveals DNA methylation profiles of integrated genome-wide and critical milk-related genes, Oncotarget, 8, 115326–115344, 2017.
Zhang, Y., Li, F., Feng, X., Yang, H., Zhu, A., Pang, J., Han, L., Zhang, T., Yao, X., and Wang, F.: Genome-wide analysis of DNA Methylation profiles on sheep ovaries associated with prolificacy using whole-genome Bisulfite sequencing, BMC Genomics, 18, 759, https://doi.org/10.1186/s12864-017-4068-9, 2017.