the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Interbreed variation in meiotic recombination rate and distribution in the domestic chicken Gallus gallus
Lyubov P. Malinovskaya
Katerina V. Tishakova
Natalia A. Volkova
Anna A. Torgasheva
Yakov A. Tsepilov
Pavel M. Borodin
The efficiency of natural and artificial selection is critically dependent on the recombination rate. However, interbreed and individual variation in recombination rate in poultry remains unknown. Conventional methods of analysis of recombination such as genetic linkage analysis, sperm genotyping and chiasma count at lampbrush chromosomes are expensive and time-consuming. In this study, we analyzed the number and distribution of recombination nodules in spermatocytes of the roosters of six chicken breeds using immunolocalization of key proteins involved in chromosome pairing and recombination. We revealed significant effects of breed (R2=0.17; p<0.001) and individual (R2=0.28; p<0.001) on variation in the number of recombination nodules. Both interbreed and individual variations in recombination rate were almost entirely determined by variation in recombination density on macrochromosomes, because almost all microchromosomes in each breed had one recombination nodule. Despite interbreed differences in the density of recombination nodules, the patterns of their distribution along homologous chromosomes were similar. The breeds examined in this study showed a correspondence between the age of the breed and its recombination rate. Those with high recombination rates (Pervomai, Russian White and Brahma) are relatively young breeds created by crossing several local breeds. The breeds displaying low recombination rate are ancient local breeds: Cochin (Indo-China), Brown Leghorn (Tuscany, Italy) and Russian Crested (the European part of Russia).
- Article
(759 KB) - Full-text XML
- BibTeX
- EndNote
Recombination makes a substantial contribution to genetic and phenotypic variability. Therefore, the efficiency of natural and artificial selection is critically dependent on the recombination rate (Battagin et al., 2016; Gonen et al., 2017). It has been shown that populations with higher recombination rate demonstrate a stronger response to selection (Gorlov et al., 1992; Korol and Iliadi, 1994; Otto and Barton, 2001). Variation in recombination rate between species (Dumont and Payseur, 2008, 2011a; Smukowski and Noor, 2011) and individuals (Broman et al., 1998; Koehler et al., 2002) has been extensively studied in mammals but rather poorly in birds.
Data obtained to date indicate much higher and less variable recombination rate in birds compared to mammals (Semenov et al., 2018). High recombination rate is determined, at least partly, by a higher proportion (about two-thirds) of microchromosomes carrying at least one obligatory chiasma. Low interspecies variation in recombination rate is usually explained by a low variation in chromosome number. In most birds, it is almost the same (2n=78–80) (Griffin et al., 2007). However, Malinovskaya et al. (2018) demonstrated that the bird species that differ substantially in chromosome number may have the same recombination rate. On the other hand, the species with the same chromosome number show significant differences in recombination rate (Calderon and Pigozzi, 2006; Semenov et al., 2018).
Variation in recombination rate, which is not explained by variation in chromosome number, is especially interesting for animal breeding. Studies in mammals revealed substantial additive genetic components of such variation. The heritability of recombination rate was estimated as 0.30 in humans, 0.22 to 0.26 in cattle and 0.15 in sheep (Kong et al., 2004; Sandor et al., 2012; Johnston et al., 2016). Genes controlling global and local variation in recombination rate in humans and mice have been identified and mapped (Fledel-Alon et al., 2011; Dumont and Payseur, 2011b).
Recombination studies in poultry are less advanced. Female-, male- and sex-averaged genetic maps of chicken chromosomes based on the results of genetic recombination analysis of F1 hybrids between the red jungle fowl and the White Leghorn were compiled (Groenen et al., 2009). Using cytological methods, Rodionov et al. (1992) and Pigozzi (2001) obtained precise estimates of the rate and distribution of recombination events along individual chromosomes of female chickens. However, interbreed and individual variation in recombination rate and distribution in chicken remains unknown. Meanwhile, this information is important for understanding the effect of selection for productivity traits on the recombination rate. It may also predict the efficiency of selection for recombination.
Conventional methods used for analysis of recombination such as genetic linkage analysis, sperm genotyping and chiasma count on lampbrush chromosomes are expensive and time-consuming. Immunolocalization of MLH1, a mismatch repair protein of mature recombination nodules at the synaptonemal complexes (SCs), has proved to produce reliable estimates of the overall recombination frequency and of the distribution of recombination events along individual chromosomes. It was successfully used for studying recombination in many vertebrates (Anderson et al., 1999; Froenicke et al., 2002; Pigozzi, 2016; Segura et al., 2013).
In this paper, we used MLH1 immunolocalization to examine variation in meiotic recombination rate and distribution in domestic chicken. We chose six breeds for our comparative study. They differed in the traits they have been selected for. Cochin and Brahma are selected for meat, Brown Leghorn and Russian White for eggs, and Russian White and Pervomai for both traits. The breeds also differ in their breeding history. Three of them (Cochin, Brown Leghorn and Russian Crested) are ancient native breeds, while the other three (Pervomai, Russian White and Brahma) are relatively modern, created by crossing several native breeds.
We addressed several questions. What is the relative contribution of interbreed and individual variations in the overall variability of recombination rate? Do the interbreed differences in recombination rate depend on the particular trait or do they depend on how long each of the particular breeds has been in existence? Does the distribution of recombination events along each macrochromosome follow a stable pattern or does it vary from one breed to the next? Answering these questions might shed light on the relationships between recombination rate and selection for economical traits in poultry.
2.1 Animals
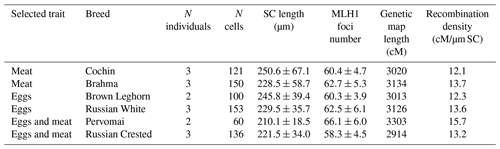
Sixteen adult five-month-old roosters of six different breeds were used in this study (Table 1). The roosters were raised and maintained at the poultry farm of the Federal Scientific Centre for Animal Husbandry under conventional conditions. Maintenance, handling and euthanasia of animals were carried out in accordance with the approved national guidelines for the care and use of laboratory animals. All experiments were approved by the Ethics Committee on Animal Care and Use at the Institute of Cytology and Genetics of the Siberian Department of the Russian Academy of Sciences (approval no. 35 of 26 October 2016).
2.2 Synaptonemal complex spreading and immunostaining
Chromosome spreads were prepared from the right testes by a drying-down method (Peters et al., 1997). Immunostaining was performed according to Anderson et al. (1999) using rabbit polyclonal anti-SYCP3 (1:500; Abcam, Cambridge, UK), mouse monoclonal anti-MLH1 (1:30; Abcam, Cambridge, UK) and human anticentromere (ACA) (1:70; Antibodies Inc., Davis, USA) primary antibodies. Primary antibody incubations were performed overnight in a humid chamber at 37 ∘C. The secondary antibodies used were Cy3-conjugated goat anti-rabbit (1:500; Jackson ImmunoResearch, West Grove, USA), fluorescein isothiocyanate (FITC)-conjugated goat anti-mouse (1:30; Jackson ImmunoResearch, West Grove, USA) and aminomethylcoumarin (AMCA)-conjugated donkey anti-human (1:40; Jackson ImmunoResearch, West Grove, USA) antibodies. Antibodies were diluted in PBT (3 % bovine serum albumin and 0.05 % Tween 20 in PBS). A solution of 10 % PBT was used for blocking non-specific binding of antibodies. Secondary antibody incubations were carried out for 1 h at 37 ∘C. Slides were mounted in Vectashield antifade mounting medium (Vector Laboratories, Burlingame, CA, USA) to reduce fluorescence fading. The preparations were visualized with an Axioplan 2 microscope (Carl Zeiss, Germany) equipped with a CCD camera (CV M300, JAI Corporation, Yokohama, Japan), CHROMA filter sets and ISIS4 image-processing package (MetaSystems GmbH, Altlußheim, Germany).
2.3 Image analysis
MLH1 signals were scored only if they were localized on synaptonemal complexes (SCs). The length of each SC and the total SC length in each spermatocyte were measured in micrometers using MicroMeasure 3.3 software (Reeves, 2001). The positions of centromeres and of MLH1 foci relative to the centromere were recorded. We identified individual SCs by their relative lengths and centromeric indices. To visualize the pattern of MLH1 foci distribution along each macrochromosome, we divided average length of the seven largest macroSCs by intervals and plotted the proportion of MLH1 foci located within each interval. To visualize the dependence of the recombination pattern on the chromosome size, we set the number of intervals for each macrochromosome proportional to the average SC length, each interval being ∼1 µm in length. We used two estimates of crossover interference: the relative distance between two neighboring MLH1 foci (the shorter the distance, the weaker the interference) and the shape parameter ν of the distribution of the distances fitted to gamma distribution. The ν values vary from 0 (no interference) to 20 (absolute interference, or only one crossover per chromosome) (Falque et al., 2009; Gauthier et al., 2011). The shape parameter ν of the distribution was estimated using CODA v.1.1 software (Gauthier et al., 2011). The STATISTICA 6.0 software package (StatSoft) was used for descriptive statistics.
The influence of interbreed and individual variations on the overall number of MLH1 foci per cell was estimated as the proportion of the variance explained by the variable (R2) by two-way ANOVA using R software (function “aov”). Differences between the breeds in the average number of MLH1 foci per each macrochromosome and in SC length were estimated by Student's t test. p<0.05 was considered to be statistically significant. Values in the text and tables are presented as means ± SD.
We analyzed the number and distribution of MLH1 foci at 28 080 completely synapsed SCs in 720 spermatocytes of 16 roosters. The rooster pachytene karyotype contained 38 autosomal SCs and a ZZ pair. We identified the seven largest macroSCs by their relative lengths and centromeric indices. SC1, SC2 and SCZZ were large metacentrics. They differed from each other in length and centromeric indices (p<0.001). SC3 and SC5 were large- and medium-sized acrocentics, while SC4 and SC6 were medium-sized submetacentics, which also differed from each other in their relative lengths and centromeric indices. The macroSCs 7–10 and all microSCs were acrocentric, with gradually decreasing chromosomal sizes (Fig. 1).
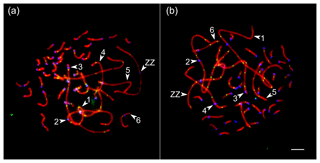
Figure 1Synaptonemal complexes (SCs) of Russian Crested (a) and Russian White (b) roosters after immunolocalization of SYCP3 (red), centromeres (blue) and MLH1 (green). Arrowheads point to the SCs of the macrochromosomes identified by their lengths and centromere indices. Bar: 5 µm.
The breeds showed a wide variation in the total length of their SCs and the number of MLH1 per cell (Table 1). To estimate the length of the recombination map in centimorgans (cM), we multiplied the average number of MLH1 foci per cell by 50 map units (one recombination event is equivalent to 50 cM). We detected a significant effect of breed (R2=0.17; p<0.001), but not of trait (p=0.12), on variation in MLH1 foci number. The effect of individual was also significant (ANOVA, R2=0.28; p<0.001). Both interbreed and individual variations in MLH1 foci number were almost entirely determined by variation in MLH1 foci density on macrochromosomes, because almost each microchromosome in each breed had one MLH1 focus.
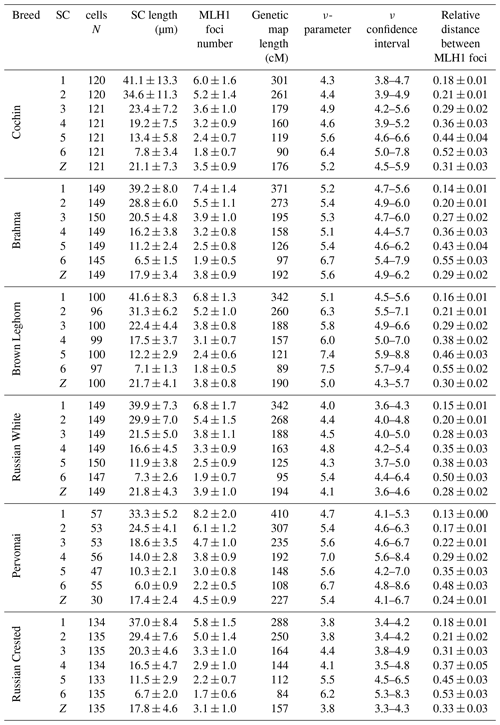
We also estimated the recombination characteristics of individual macrochromosomes: the SC length, the number of recombination nodules, the level of crossover interference (Table 2) and the pattern of MLH1 foci distribution along the SCs (Fig. 2). Table 2 shows that interbreed variation in the recombination characteristics of individual macrochromosomes was also substantial.
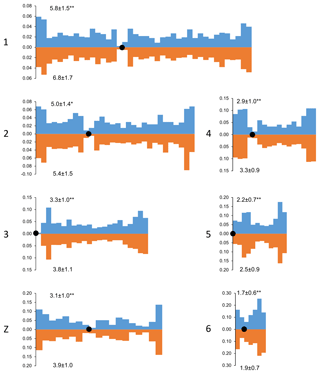
Figure 2Distribution of MLH1 foci along the macrochromosomes of Russian Crested (blue columns) and Russian White (orange columns) roosters. On the x axis: positions of the MLH1 foci on the chromosomes relative to the centromere (black circle). Each interval is the proportion of the average length of each SC and is approximately 1 µm in length. On the y axis: the proportion of MLH1 foci within each interval. The numbers to the left of the y axis stand for chromosome numbers; the numbers above and below each graph show the average number of MLH1 foci at a given chromosome of a given breed. Differences between the breeds in the average number of MLH1 foci are significant (Student's t test : p<0.01; *: p<0.05).
Despite the interbreed differences in MLH1 foci density, the patterns of their distribution along homologous chromosomes were strikingly similar (Fig. 2). We observed an increase in MLH1 foci frequency in the distal regions of all macrochromosomes. The lack of MLH1 foci in proximal regions (centromeric interference) was limited to a small pericentromeric area of about 1–2 µm in length.
Chicken macrochromosomes showed a positive correlation between SC length and number of MLH1 sites (R=0.92, p<0.001), which is typical of vertebrate chromosomes. There was a negative correlation between SC length and degree of crossover interference estimated through the relative distances between adjacent MLH1 sites (, p<0.05) and through the ν parameter of gamma distribution of the distances (, p<0.05).
In this study, we estimated the number and distribution of recombination events in the rooster genome by immunolocalization of MLH1 protein, a reliable marker of mature recombination nodules. The main advantage of this approach over chiasma count consists in the possibility to analyze a large number of both oocytes and spermatocytes, while the general estimates of recombination rate obtained by these two methods are rather close. Rodionov et al. (1992) estimated the total number of chiasmata per chicken oocyte as 59–64, and Pigozzi (2001) estimated the number of MLH1 foci per oocyte as 65.0±4.0. In our study, the number of MLH1 foci per spermatocyte varied from 58.3±4.5 to 66.1±6.0 across the chicken breeds, with the general average being 61.5±5.6.
The results of cytological and genetic analyses of recombination are in a good agreement with each other. For example, Groenen et al. (2009) estimated the male chicken genetic map length as 3145 cM based on the results of genetic linkage analysis of F1 hybrids between the red jungle fowl and the White Leghorn. The breed-averaged male chicken genetic map resulting from our study was just 1 % shorter (3087 cM). Thus, we recommend the immunolocalization of MLH1 protein as a fast, affordable and reliable method for recombination analysis in poultry.
The most important and interesting results of our study are (1) the detection of wide individual and interbreed variation in recombination rate (both overall and macrochromosome-specific) and (2) a remarkable stability of the pattern of distribution of recombination events along macrochromosomes.
The divergence of the breeds in recombination rate may have been driven by differences in breeding histories. The breeds examined showed an interesting correspondence between the age of the breed and its recombination rate. Those with high recombination rate are relatively young breeds created by crossing several local stocks. The Pervomai breed was produced by complex reproductive crossing of three crossbred breeds: White Wyandotte (derived from crosses between Brahmas and Hamburgs), Rhode Island (derived from crosses between Malays and brown Italian Leghorns) and Yurlov Crower (derived from crosses of Chinese meat chicken, gamecocks and landraces) in 1930–1960. The Russian White originated from crosses of the White Leghorn with local chickens in 1920–1940. The Brahma breed was developed in the United States in 1840–1850 by crossing the ancient Asian breeds Cochin and Chittagong. On the other hand, the breeds displaying low recombination rate are ancient local breeds: Cochin (Indo-China), Brown Leghorn (Tuscany, Italy) and Russian Crested (the European part of Russia) (Paronyan and Yurchenko, 1989; Scrivener, 2006, 2009).
We may speculate that the correspondence between recombination rate and breed's age appears to reflect a correspondence between recombination rate and genetic heterogeneity because genetic heterogeneity within each breed tends to decrease with time due to inbreeding and artificial selection (Lipinski et al., 2008; Gibbs et al., 2009). This hypothesis is consistent with the assumption that benefits of recombination (generation of new allele combinations) prevail over its costs (occurrence of deleterious mutations) as long as the population remains sufficiently heterogeneous (Kim and Stephan, 2000; Nachman, 2001; Ohta, 1999). Thus, depletion of genetic variability may lead to a decrease in recombination efficiency, which reduces selection pressure on high recombination rate. This would in turn decrease recombination rate. Further studies are needed to check these hypotheses.
The conservatism in the distribution of recombination events across the rooster macrochromosomes confirms the pattern previously described in mammals: the recombination frequency in certain regions of vertebrate chromosomes depends more upon the position of these regions on the chromosome (that is, upon the distance from the centromere and the telomere) than upon the genetic composition of these regions (Gorlov et al., 1991; Vozdova et al., 2016; Ruiz-Herrera et al., 2017).
Our results are relevant to the current discussion about the role of recombination in selection for productivity traits. Because recombination reduces linkage disequilibrium between quantitative trait loci, it is considered a means to cope with negative epistasis and to deal with the selection plateau. It has been shown with a variety of model organisms and by computational experiments under different scenarios (Comeron et al., 1999; Pál et al., 2001; Carvalho and Clark, 2002; Simmonds, 2006; Hill and Robertson, 2008) that an increase in recombination rate enhances the efficiency of selection. However, the magnitude of this effect is still the subject of a debate. A recent simulation study by Battagin et al. (2016) demonstrated that the effect will be taking place in the distant future and requires an unrealistically high increase in global recombination rate. The simulation shows that a 2-fold increase in recombination rate leads to a 12.5 % increase in cumulative selection response over 40 generations, and a 20-fold increase in recombination rate results in a 33.4 % increase in the response to selection.
In our study, the difference in recombination rate between the most contrasted breeds, Pervomai and Russian Crested, was 19 %. Moreover, the results of our study demonstrate that the interbreed differences in recombination rate have no effect on the distribution of recombination nodules. Their localization along the chicken macrochromosomes is chromosome-specific, positionally determined and conserved between the breeds. This indicates that selection of overall recombination rate in chickens is unlikely to affect the efficiency of selection for productivity within the breeds. Apparently, the selection success could be achieved by manipulation of local rather than general recombination rate. The detection and mapping of hot and cold spots, which show up as 1000-fold differences in recombination rate, and the proteins involved in their regulation could make this approach feasible (Elferink et al., 2010; Gonen et al., 2017; Singhal et al., 2015). This targeted approach might help to break up the ancestral linkage between plus and minus alleles for productivity traits.
The data sets are available upon request from the corresponding author.
PMB and NAV designed the experiment. LPM, KVT and AAT prepared and analyzed chromosome preparations. YAT performed statistical analysis. PMB, AAT, NAV and YAT wrote and edited the paper.
The authors declare that they have no conflict of interest.
This work was supported by the Russian Foundation for Basic Research (grant no. 17-29-08019) and the Ministry of Science and Higher Education via the Institute of Cytology and Genetics (grant no. 0324-2019-0042). The funding bodies play no role in the design of the study and collection, analysis and interpretation of data, and in writing the manuscript.
This research has been supported by the Russian Foundation for Basic Research (grant no. 17-29-08019) and the Ministry of Science and High Education of Russian Federation (grant no. 0324-2019-0042).
This paper was edited by Steffen Maak and reviewed by two anonymous referees.
Anderson, L. K., Reeves, A., Webb, L. M., and Ashley, T.: Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein, Genetics, 151, 1569–1579, 1999.
Battagin, M., Gorjanc, G., Faux, A. M., Johnston, S. E., and Hickey, J. M.: Effect of manipulating recombination rates on response to selection in livestock breeding programs, Genet. Sel. Evol., 48, 44, https://doi.org/10.1186/s12711-016-0221-1, 2016.
Broman, K. W., Murray, J. C., Sheffield, V. C., White, R. L., and Weber, J. L.: Comprehensive human genetic maps: individual and sex-specific variation in recombination, Am. J. Hum. Genet., 63, 861–869, 1998.
Calderon, P. L. and Pigozzi, M. I.: MLH1-focus mapping in birds shows equal recombination between sexes and diversity of crossover patterns, Chromosom. Res., 14, 605–612, 2006.
Carvalho, A. B. and Clark, A. G.: Intron size and natural selection, Nature, 401, 344, 2002.
Comeron, J. M., Kreitman, M., and Aguade M.: Natural selection on synonymous sites is correlated with gene length and recombination in Drosophila, Genetics, 151, 239–249, 1999.
Dumont, B. L. and Payseur, B. A.: Evolution of the genomic rate of recombination in mammals, Evolution, 62, 276–294, 2008.
Dumont, B. L. and Payseur, B. A.: Evolution of the genomic recombination rate in murid rodents, Genetics, 187, 643–657, 2011a.
Dumont, B. L. and Payseur, B. A.: Genetic Analysis of Genome-Scale Recombination Rate Evolution in House Mice, PLoS Genet., 7, e1002116, https://doi.org/10.1371/journal.pgen.1002116, 2011b.
Elferink, M. G., van As, P., Veenendaal, T., Crooijmans, R. P. M. A., and Groenen, M. A. M.: Regional differences in recombination hotspots between two chicken populations, BMC Genet., 11, 11, https://doi.org/10.1186/1471-2156-11-11, 2010.
Falque, M., Anderson, L. K., Stack, S. M., Gauthier, F., and Martin, O. C.: Two Types of Meiotic Crossovers Coexist in Maize, Plant Cell, 21, 3915–3925, 2009.
Fledel-Alon, A., Leffler, E. M., Guan, Y., Stephens, M., Coop, G., and Przeworski, M.: Variation in human recombination rates and its genetic determinants, PLoS One, 6, e20321, https://doi.org/10.1371/journal.pone.0020321, 2011.
Froenicke, L., Anderson, L. K., Wienberg, J., and Ashley, T.: Male mouse recombination maps for each autosome identified by chromosome painting, Am. J. Hum. Genet., 71, 1353–1368, 2002.
Gauthier, F., Martin, O. C. O., Falque, M., Jones, G., Franklin, F., Anderson, L., Stack, S., Sturtevant, A., Berchowitz, L., Francis, K., Bey, A., Copenhaver, G., Lhuissier, F., Offenberg, H., Wittich, P., Vischer, N., Heyting, C., Copenhaver, G., Housworth, E., Stahl, F., Housworth, E., Stahl, F., Falque, M., Anderson, L., Stack, S., Gauthier, F., Martin, O. C. O., Kleckner, N., Zickler, D., Jones, G., Dekker, J., Padmore, R., Henle, J., Hutchinson, J., McPeek, M., Speed, T., Zhao, H., McPeek, M., Speed, T., Viswanath, L., Housworth, E., Housworth, E., Stahl, F., Saintenac, C., Falque, M., Martin, O. C. O., Paux, E., Feuillet, C., Sourdille, P., Broman, K., Rowe, L., Churchill, G., Paigen, K., Anderson, L., Doyle, G., Brigham, B., Carter, J., Hooker, K., Lai, A., Rice, M., Stack, S., Foss, E., Lande, R., Stahl, F., and Steinberg, C.: CODA (crossover distribution analyzer): quantitative characterization of crossover position patterns along chromosomes, BMC Bioinform., 12, 27, https://doi.org/10.1186/1471-2105-12-27, 2011.
Gibbs, R. A., Taylor, J. F., Van Tassell, C. P., Barendse, W., Eversole, K. A., Gill, C. A., Green, R. D., Hamernik, D. L., Kappes, S. M., Lien, S., Matukumalli, L. K., McEwan, J. C., Nazareth, L. V., Schnabel, R. D., Weinstock, G. M., Wheeler, D. A., Ajmone-Marsan, P., Barends, W., Boettcher, P. J., Caetano, A. R., Garcia, J. F., OlivierHanotte, Mariani, P., Skow, L. C., Sonstegard, T. S., Williams, J. L., Diallo, B., Hailemariam, L., Hanotte, O., Martinez, M. L., Morris, C. A., Silva, L. O. C., Spelman, R. J., Mulatu, W., Zhao, K., Abbey, C. A., Agaba, M., Araujo, F. R., Bunch, R. J., Burton, J., Gorni, C., Harrison, B. E., Luff, B., Machado, M. A., Mwakaya, J., Plastow, G., Sim, W., Smith, T., Thomas, M. B., Valentini, A., Williams, P., Womack, J., Woolliams, J. A., Liu, Y., Qin, X., Worley, K. C., Gao, C., Jiang, H., Moore, S. S., Ren, Y., Song, X. Z., Bustamante, C. D., Hernandez, R. D., Muzny, D. M., Patil, S., Lucas, A. S., Fu, Q., Kent, M. P., Vega, R., Matukumalli, A., McWilliam, S., Sclep, G., Bryc, K., Choi, J., Gao, H., Grefenstette, J. J., Murdoch, B., Stella, A., Villa-Angulo, R., Wright, M., Aerts, J., Jann, O., Negrini, R., Goddard, M. E., Hayes, B. J., Bradley, D. G., Da Silva, M. B., Lau, L. P. L., Liu, G. E., Lynn, D. J., Panzitta, F., and Dodds, K. G.: Genome-wide survey of SNP variation uncovers the genetic structure of cattle breeds, Science, 324, 528–532, 2009.
Gonen, S., Battagin, M., Johnston, S. E., Gorjanc, G., and Hickey, J. M.: The potential of shifting recombination hotspots to increase genetic gain in livestock breeding, Genet. Sel. Evol., 49, 55, https://doi.org/10.1186/s12711-017-0330-5, 2017.
Gorlov, I. P., Ladygina, T. Y., Serov, O. L., and Borodin, P. M.: Positional control of chiasma distribution in the house mouse. Chiasma distribution in mice homozygous and heterozygous for an inversion in chromosome 1, Heredity, 66, 453–458, 1991.
Gorlov, I. P., Schuler, L., Bunger, L., and Borodin, P.: Chiasma frequency in strains of mice selected for litter size and for high body weight, TAG Theor. Appl. Genet., 84, 640–642, 1992.
Griffin, D. K., Robertson, L. B. W., Tempest, H. G., and Skinner, B. M.: The evolution of the avian genome as revealed by comparative molecular cytogenetics, Cytogenet. Genome Res., 117, 64–77, 2007.
Groenen, M. A. M., Wahlberg, P., Foglio, M., Cheng, H. H., Megens, H. J., Crooijmans, R. P. M. A. R. P., Besnier, F., Lathrop, M., Muir, W. M., Wong, G. K. S., Gut, I., and Andersson, L.: A high-density SNP-based linkage map of the chicken genome reveals sequence features correlated with recombination rate, Genome Res., 19, 510–519, 2009.
Hill, W. G. and Robertson, A.: The effect of linkage on limits to artificial selection, Genet. Res. (Camb), 8, 269–294, 2008.
Johnston, S. E., Bérénos, C., Slate, J., and Pemberton, J. M.: Conserved genetic architecture underlying individual recombination rate variation in a wild population of soay sheep (Ovis aries), Genetics, 203, 583–598, 2016.
Kim, Y. and Stephan, W.: Joint Effects of Genetic Hitchhiking and Background Selection on Neutral Variation, Genetics, 155, 1415–1427, 2000.
Koehler, K. E., Cherry, J. P., Lynn, A., Hunt, P. A., and Hassold, T. J.: Genetic control of mammalian meiotic recombination. I. Variation in exchange frequencies among males from inbred mouse strains, Genetics, 162, 297–306, 2002.
Kong, A., Barnard, J., Gudbjartsson, D. F., Thorleifsson, G., Jonsdottir, G., Sigurdardottir, S., Richardsson, B., Jonsdottir, J., Thorgeirsson, T., Frigge, M. L., Lamb, N. E., Sherman, S., Gulcher, J. R., and Stefansson, K.: Recombination rate and reproductive success in humans, Nat. Genet., 36, 1203–1206, 2004.
Korol, A. B. and Iliadi, K. G.: Increased recombination frequencies resulting from directional selection for geotaxis in Drosophila, Heredity, 72, 64–68, 1994.
Lipinski, M. J., Froenicke, L., Baysac, K. C., Billings, N. C., Leutenegger, C. M., Levy, A. M., Longeri, M., Niini, T., Ozpinar, H., Slater, M. R., Pedersen, N. C., and Lyons, L. A.: The ascent of cat breeds: Genetic evaluations of breeds and worldwide random-bred populations, Genomics, 91, 12–21, 2008.
Malinovskaya, L., Shnaider, E., Borodin, P., and Torgasheva, A.: Karyotypes and recombination patterns of the Common Swift (Apus apus Linnaeus, 1758) and Eurasian Hobby (Falco subbuteo Linnaeus, 1758), Avian Res., 9, 4, https://doi.org/10.1186/s40657-018-0096-7, 2018.
Nachman, M. W.: Single nucleotide polymorphisms and recombination rate in humans, Trends Genet., 17, 481–485, 2001.
Ohta, T.: A note on the correlation between heterozygosity and recombination rate, Genes Genet. Syst., 74, 209-210, 1999.
Otto, S. P. and Barton, N. H.: Selection for recombination in small populations, Evolution, 55, 1921–1931, 2001.
Pál, C., Papp, B., and Hurst, L. D.: Does the recombination rate affect the efficiency of purifying selection? The yeast genome provides a partial answer, Mol. Biol. Evol., 18, 2323–2326, 2001.
Paronyan, I. A. and Yurchenko, O. P.: Domestic fowl, in: Animal genetic resources of the USSR, edited by: Dmitriev, N. G. and Ernst, L. K., Food and Agriculture Organization of the United Nations, Rome, 437–469, 1989.
Peters, A. H., Plug, A. W., van Vugt, M. J., and de Boer, P.: A drying-down technique for the spreading of mammalian meiocytes from the male and female germline, Chromosome Res., 5, 66–68, 1997.
Pigozzi, M. I.: Distribution of MLH1 foci on the synaptonemal complexes of chicken oocytes, Cytogenet. Cell Genet., 95, 129–133, 2001.
Pigozzi, M. I.: The chromosomes of birds during meiosis, Cytogenet. Genome Res., 150, 128–138, 2016.
Reeves, A.: MicroMeasure: a new computer program for the collection and analysis of cytogenetic data, Genome, 44, 439–443, 2001.
Rodionov, A. V, Miakoshina, Y. A., Chelysheva, L. A., Solovei, I. V., and Gaginskaya, E. P.: Chiasmata at lampbrush chromosomes of Gallus gallus domesticus: cytogenetic study of recombination frequency and linkage group length, Genetika, 28, 53–63, 1992.
Ruiz-Herrera, A., Vozdova, M., Fernández, J., Sebestova, H., Capilla, L., Frohlich, J., Vara, C., Hernández-Marsal, A., Sipek, J., Robinson, T. J., and Rubes, J.: Recombination correlates with synaptonemal complex length and chromatin loop size in bovids – insights into mammalian meiotic chromosomal organization, Chromosoma, 126, 615–631, 2017.
Sandor, C., Li, W., Coppieters, W., Druet, T., Charlier, C., and Georges, M.: Genetic variants in REC8, RNF212, and PRDM9 influence male recombination in cattle, PLoS Genet., 8, e1002854, https://doi.org/10.1371/journal.pgen.1002854, 2012.
Scrivener, D.: Rare poultry breeds, The Crowood Press Ltd, Marlborough, 2006.
Scrivener, D.: Popular poultry breeds, The Crowood Press Ltd, Marlborough, 2009.
Segura, J., Ferretti, L., Ramos-Onsins, S., Capilla, L., Farré, M., Reis, F., Oliver-Bonet, M., Fernández-Bellón, H., Garcia, F., Garcia-Caldés, M., Robinson, T. J., and Ruiz-Herrera, A.: Evolution of recombination in eutherian mammals: insights into mechanisms that affect recombination rates and crossover interference, P. Roy. Soc. B, 280, 20131945, https://doi.org/10.1098/rspb.2013.1945, 2013.
Semenov, G., Basheva, E. A., Borodin, P. M., and Torgasheva, A. A.: High rate of meiotic recombination and its implications for intricate speciation patterns in the white wagtail (Motacilla alba), Biol. J. Linn. Soc., 125, 600–612, 2018.
Simmonds, P.: Recombination and Selection in the Evolution of Picornaviruses and Other Mammalian Positive-Stranded RNA Viruses, J. Virol., 80, 11124–11140, 2006.
Singhal, S., Leffler, E. M., Sannareddy, K., Turner, I., Venn, O., Hooper, D. M., Strand, A. I., Li, Q. Y., Raney, B., Balakrishnan, C. N., Griffith, S. C., McVean, G., and Przeworski, M.: Stable recombination hotspots in birds, Science, 350, 928–932, 2015.
Smukowski, C. S. and Noor, M. A. F.: Recombination rate variation in closely related species, Heredity, 107, 496–508, 2011.
Vozdova, M., Ruiz-Herrera, A., Fernandez, J., Cernohorska, H., Frohlich, J., Sebestova, H., Kubickova, S., and Rubes, J.: Meiotic behaviour of evolutionary sex-autosome translocations in Bovidae, Chromosom. Res., 24, 325–338, 2016.