the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
The novel 22 bp insertion mutation in a promoter region of the PITX2 gene is associated with litter size and growth traits in goats
Hailong Yan
Enhui Jiang
Haijing Zhu
Linyong Hu
Jinwang Liu
Lei Qu
The paired-like homeodomain 2 (PITX2) gene plays a critical role in regulating development, reproduction, and growth traits in ruminants. Hence, the objective of this study was to explore the polymorphisms of this gene and to evaluate their associations with quantitative traits. Herein, a novel insertion in the promoter region of the PITX2 gene was reported in Shaanbei white cashmere (SBWC) goats (n=1012). The genotype distributions between mothers of single-kid and multi-kid groups within SBWC goats were significantly different (P<0.01), implying that this indel mutation might affect the litter size. Furthermore, association analysis found that this indel mutation was significantly associated with litter size (P=0.001). Individuals with genotype DD had a significantly smaller litter size than those with other genotypes (P<0.01). Besides, this indel was significantly associated with the body length (P=0.042) and the chest width (P=0.031). Especially, the individuals with genotype DD had a significantly lower body length than those with genotype II (P<0.05), which was consistent with the trend in litter size. These findings suggested that the new 22 bp indel mutation within the PITX2 gene is significantly associated with litter size and growth traits; this can be utilized as a functional molecular marker in goat breeding.
- Article
(302 KB) - Full-text XML
- Companion paper
- BibTeX
- EndNote
The PITX2 gene was originally identified as the gene associated with human Rieger syndrome, an autosomal dominant disease (Hassed et al., 2017). Subsequently, this gene's additional functions were studied widely. The available studies have shown that the PITX2 gene is important for asymmetric development of organs in animals (Welsh et al., 2013; Syeda et al., 2017). Thus, this gene is essential for the normal physiological functioning in animals (S. H. Zhang et al., 2018). In addition to Rieger syndrome, mutation or misregulation of PITX2 can also cause other diseases such as atrial fibrillation and congenital glaucoma (Yin et al., 2014; Zhou et al., 2015; Chen et al., 2016; Li et al., 2016). Furthermore, reports have also revealed that the PITX2 gene is important in muscle formation and development. In the mouse model, PITX2 together with similar upstream regulators, PAX3 and PAX7, participates in gene regulatory networks at different skeletal myogenesis sites (Margaret and Peter, 2014). Likewise, the PITX2 gene can modulate fetal and adult myogenesis by controlling redox conditions and regulating MyoD, respectively (L'Honoré et al., 2010; Hernandez et al., 2017).
In addition to muscle development, evidence shows that the PITX2 gene plays irreplaceable roles in signal pathways such as Wnt∕β-catenin and POU1F1–PROP1. (1) In Wnt∕β-catenin signal pathway, the PITX2 gene not only activates the canonical pathway by itself and by inducing Wnt ligands but also limits further Wnt activation by down-regulating frizzled receptors. The PITX2 gene can also mediate mRNA stabilization in the Wnt∕β-catenin pathway (Basu et al., 2013). Essential role played by Wnt∕β-catenin signal pathway in regulation of progenitor cell fate, development decisions, proliferation during embryonic development, and adult tissue homeostasis is known (Maryam et al., 2018). Consequently, the PITX2 gene plays a role in diverse functions through Wnt∕β-catenin signal pathway. (2) In POU1F1–PROP1 signal pathway, the PITX2 gene regulates transcription and expression of genes such as POU1F1, LHX3, and PROP1, which influence the secretion of growth hormone (GH), prolactin (PRL), etc. (Lan et al., 2007; Zhao et al., 2017). These hormones are essential for growth, development, and reproduction of animals; thus, we speculate that the PITX2 gene may affect production and reproduction in animals.
In terms of reproduction, previous studies have shown that the PITX2 gene also regulates twinning (Angela et al., 2014), owing to PITX2 gene function in embryonic axis formation, and influences on embryo polarity. Therefore, PITX2 is a potential candidate gene for maker-assisted selection (MAS) in breeding and genetics. Currently, there are few studies about this gene and its effects on prolificacy and growth traits in goats.
Shaanbei white cashmere (SBWC) goat is a breed developed by crossing a Liaoning cashmere goat (male parent) and a Shaanbei black goat (female parent) and is characterized by resistance to rough feeding, cold weather, wind and disease; these goats are widely raised in northern Shaanxi province (Yang et al., 2017; Yan et al., 2018; Cui et al., 2018a). However, there are some problems in the breeding process such as thickening of cashmere fiber, decrease of the cashmere yield, and production of single kid, all of which reduce the economic benefit of cashmere goats. This study was conducted to detect possible novel polymorphisms within the PITX2 gene and to evaluate their effects on litter size and growth traits in SBWC goats.
All experimental procedures were approved by the institutional animal care and use committee (IACUC) of Northwest A&F University (NWAFU). The use of experimental animals was in accordance with the local animal welfare laws, guidelines, and policies.
2.1 DNA samples and related data collection
Ear tissue samples were obtained from 1012 unrelated female SBWC goats (2–5 years old), which were reared on the different farms, under similar managemental and feeding programs, and under normal conditions, in central Yulin, Shaanxi, China (Wang et al., 2017; Yang et al., 2017; Cui et al., 2018a; Yan et al., 2018). The feeding programs were the following: all ewes were mated with rams at a ratio of 18–24 to 1. All the kids were continuously kept with their dams up to weaning at the age of 3 months (Wang et al., 2018). Furthermore, litter size (LS) at the first birth as well as growth traits including body height (BH), body length (BL), chest width (CW), chest circumference (CC), and chest depth (CD) were measured and recorded (Wang et al., 2017; Yang et al., 2017; Zhao et al., 2017; Yan et al., 2018).
2.2 Genomic DNA isolation and DNA pool construction
All genomic DNA samples isolated from ear tissues, using standard salt–chloroform extraction method (Aljanabi et al., 1997; Sambrook and Russell, 2001; Lan et al., 2007; Zhou et al., 2016; Zhang et al., 2017), were quantified by NanoDrop 1000 (Thermo Scientific, Waltham, MA, USA) and then diluted to 50 ng/μL as working solutions (Chen et al., 2016). Fifty DNA samples were randomly selected to construct genomic DNA pool for exploring genetic variation in the PITX2 gene (Jin et al., 2016).
2.3 Primer design and PCR amplifications
In this work, two indel mutations were obtained from NCBI, which were named rs670348258 (in promoter region) and rs655548222 (in intron). Hence, a total of four primer pairs were designed using Primer Premier 5.0 software (Primer, Canada), based on the reference sequence (GenBank accession number NC_030813.1) (Table 1). Among them, primer pair P1 was used to detect the variations in rs670348258, and the others were used for those in rs655548222 (Table 1). All primer pairs were synthesized by Sangon Biotech (Xian, Shaanxi, China). Polymerase chain reaction (PCR) was performed in 12.5 µL reaction mixture, containing 1.5 µL (10 ng∕µL) of genomic DNA, 0.5 µL of each primer, 6.0 µL 2×Taq Master Mix, and 4.0 µL ddH2O. Two thermal cycling programs were used in this study. The first program involved pre-denaturation for 5 min at 95 ∘C followed by 34 cycles of denaturation at 94 ∘C for 30 s; annealing for 30 s at 52 ∘C, elongation for 15 s at 72∘C; and a final extension at 72 ∘C for 10 min. The second program, touch-down PCR (TD-PCR), included pre-denaturation for 5 min at 95 ∘C; followed by 18 cycles of denaturation at 94 ∘C for 30 s; annealing for 30 s at 68 ∘C (with a decrease of 1 ∘C per cycle); elongation for 15 s at 72 ∘C; and final extension at 72 ∘C for 10 min (Cui et al., 2018a; Y. H. Zhang et al., 2018). Finally, 4.0 µL PCR products were directly assayed by electrophoresis on 3.0 % agarose gels stained with ethidium bromide.
2.4 Statistical analysis
Genotypic and allelic frequencies were calculated manually. Hardy–Weinberg equilibrium (HWE) analysis was performed using the SHEsis program (http://analysis.bio-x.cn, last access: 6 August 2018) (Yan et al., 2018). The parameters of population genetics are the following: heterozygosity (He), homozygosity (Ho), effective allele number (Ne) and polymorphism information content (PIC) were calculated by PopGene version 1.3.1 (Molecular Biology and Biotechnology Centre, University of Alberta, Edmonton, AB, Canada) (Wang et al., 2018).
Distribution differences, for genotypic frequencies, between the mothers of a single kid and multiple kids were analyzed by χ2 test (Li et al., 2018). The association of indel polymorphism with several traits was measured. These associations were analyzed when the data agreed with the characteristics of normality and homogeneity of variance (ANOVA). The non-parametric Tamhane test was also carried out. ANOVA applied a general linear model (GLM) and the reduced linear model as follows:
where Yi is the trait measured for each animal, u the mean for each trait, Gi the fixed effect of genotype or combined genotype, and e the random error (Wang et al., 2018; Yang et al., 2017; Cui et al., 2018a; Yan et al., 2018).
Besides, genetic effects of different genotypes on litter size were also evaluated under different genetic models: codominant, dominant, recessive, and overdominant. The Akaike information criterion (AIC) was used to confirm which genetic effect model was used for the data (EI-Bayomi et al., 2018). The model with the smallest AIC value was probably the best fit based on the evaluation results. All the above analyses were carried out using SPSS software (version 18.0, IBM, USA). The results are expressed as means±SE. Differences between the means were considered significant at P<0.05 (Ren et al., 2017).
3.1 Identification of a novel 22 bp indel variation
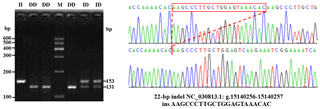
A novel 22 bp insertion was uncovered, using DNA pool sequencing (Pak et al., 2002), 967 bp upstream from the first exon of the goat PITX2 gene, and was described as NC_030813.1: g.15140256-15140257 ins AAGCCCTTGCTGGAGTAAACAC (Fig. 1). The novel indel sequence was completely different from the sequence registered as NCBI-dbSNP (AAACACAAGCCCTTGCTGGAGT), and the current position of the indel is 6 bp behind the predicted indel mutation provided by NCBI. Electrophoretic analysis showed that there were three genotypes (Fig. 1).
3.2 Genetic parameter analysis of 22 bp indel
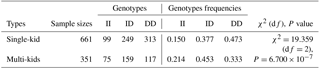
Frequency distributions of the three genotypes and two alleles are summarized in Table 2, in addition to gene homozygosity (HO), heterozygosity (He), effective allele number (Ne), and the polymorphism information content (PIC). The frequency of the DD genotype (0.425) was the highest in the studied population of SBWC goat. SBWC goat population was not at HWE (P<0.05). The value of PIC (0.358) showed that it had medium genetic diversity () at the 22 bp indel locus.
3.3 Intra-group analysis of single-kid and multi-kid individuals in SBWC goats
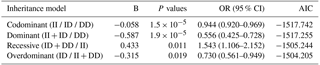
The independent intra-group χ2 test of 1012 female, goats (Table 3) showed that litter size was significantly correlated with different genotypic frequencies (χ2=19.359, df=2, ).
3.4 Association analysis between indel genotypes and litter size and growth traits in SBWC goats
The association between 22 bp indel in the PITX2 gene and litter size is shown in Table 4. Individuals with genotype DD had a significantly smaller litter size than those with other genotypes (P<0.01). Likewise, the association of the indel and some growth traits was shown in Table 4. The analysis showed that this indel was significantly associated with the body length (P=0.045) and the chest width (P=0.031). The individuals with genotype DD had a significantly lower body length than the individuals with genotype II (P<0.05), which was consistent with the trend observed in litter size. However, the individuals with genotype DD had a significantly higher chest width than the individuals with genotype II (P<0.05).
Table 4The relationship between three genotypes of the PITX2 indel variants and traits in Shaanbei white cashmere (SBWC) goats.
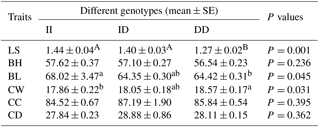
Note: BH, body height; BL, body length; CC, chest circumference; CD, chest depth; CW, chest width; LS, litter size. The value with different letters (a and b) within the same row differ significantly at P<0.05. The value with different letters (A and B) within the same row differ significantly at P<0.01.
3.5 Genetic effects of different genotypes on litter size under different models
According to the EI-Bayomi description (EI-Bayomi et al., 2018), the logistic regression analysis revealed a significant association between the insertion mutation and litter size in the codominant model (, dominant model (, recessive (P=0.011), and overdominant model (P=0.019) (Table 5). In addition, codominant model had the smallest AIC value ().
Indel mutations of the critical genes such as SPEF2 gene (Chen et al., 2016), GHR gene (Yang et al., 2017), GDF9 gene (Wang et al. 2017), LHX4 gene (Yan et al., 2018), KDM5B gene (Cui et al., 2018b), KDM6A gene (Cui et al., 2018a), and CSN1S1 gene (Wang et al., 2018) are widely known to influence production traits in animals. Several SNPs in introns and exons within the PITX2 gene have been reported in goats, but to the best of our knowledge indel polymorphisms of this gene have not been reported until now. According to previous studies, the PITX2 gene has a significant effect on milk production traits in dairy goats (Zhao et al., 2013). In this study a novel indel within the promoter of the PITX2 gene in goats was explored. To explore the correlation between the 22 bp indel and some economic traits, a large sample of goats (n=1012) was analyzed. This indel locus showed intermediate polymorphism as shown by the PIC values (). The tested SBWC population was not in accordance with Hardy–Weinberg equilibrium, suggesting that populations were not in dynamic equilibrium probably due to artificial selection, migration, and genetic drift (Zhang et al., 2015).
The influence of the indel on litter size was analyzed in two steps. First, intra-group χ2 test was utilized to analyze whether the litter size was significantly correlated with different genotypes. Results indicated that the genotypic distribution is associated with litter size because, multi-kid-mother individuals showed a higher frequency of II and ID genotypes than single-kid-mother individuals. Furthermore, association analysis showed that it was not only associated significantly with the litter size (P=0.001) but also with the body length (P=0.045) and the chest width (P=0.031). Together, the individuals with II showed better performance in litter size and body length, which was consistent with our previous reports in the same breed (Wang et al., 2017; Yang et al., 2017; Yan et al., 2018). Interestingly, the logistic regression analysis uncovered a significant association between the insertion mutation and litter size in different models. The best fit model with smallest AIC value was the codominant model.
The significant effects of the 22 bp indel on reproductive traits might be explained by the function of the PITX2 gene: first, as a member of POU1F1–PROP1 pathway, the PITX2 gene regulates transcription and expression of other genes such as POU1F1, LHX3, PROP1, GH, and PRL (Zhang et al., 2015; Bertko et al., 2017; Zhao et al., 2017). Second, the PITX2 gene regulates Wnt∕β-catenin pathway, affecting embryo implantation (Yusuke et al., 2006; Basu et al., 2013; Yeonsoo et al., 2015); therefore, this might be linked to litter size. Third, the PITX2 gene also regulates multiplets by influencing embryo polarity and axis formation (Angela et al., 2014). For example, in the chick embryo, the PITX2 gene is the direct regulator of Vg1 expression, and localized expression of Vg1 gene is responsible for formation of embryo poles (Angela et al., 2014). Deletion of Vg1 gene in zebra-fish embryo was reported to cause reproductive failure (Tessa et al., 2017). Finally, our analysis of the goat PITX2 gene via the Promoter 2.0 Prediction Server showed the location of the indel. In the sequence of I allele, the indel was located 967 bp upstream from the first exon in the promoter region, which includes the 945–1045 bp region upstream from the first exon; however, in the sequence of D allele, the indel was not located in a promoter region (545–645 bp upstream from the first exon). This indicated that the indel may influence the mechanism of the goat PITX2 promoter and the hypothesis should be verified by more studies. It is well known that promoters are essential in regulating the level of gene expression and controlling the expression in a spatial and temporal manner (Qiu et al., 2015). Therefore, the indel locus in the promoter region can influence gene expression and, consequently, certain economic traits in many species such as porcine (Cieslak et al., 2013) and cattle (Jin et al., 2016).
Additionally, the significant effect of the 22 bp indel on growth traits can also be explained by the function of the PITX2 gene: the PITX2 gene is important in muscle formation and development, and it can influence secretion of hormones such as GH and PRL. Further, the dominant genotypes of some key genes in SBWC goats can improve not only the physical traits but also the litter size. For example, Wang et al. (2017) found that there was a significant relationship between the litter size and body height in the individuals with same genotypes while exploring a 12 bp indel polymorphism within the goat GDF9 gene. Yang et al. (2017) reported that the dominant genotype of the 14 bp duplicated indel within the GHR gene simultaneously and significantly affected litter size and growth traits. Yan et al. (2018) found that a 12 bp deletion within the goat LHX4 gene significantly affected litter size. Variants of LHX4 gene were also associated with growth traits (Ren et al., 2010). Therefore, all of the evidence indicates that different alleles have a similar potential advantage in marker-assisted selection (MAS).
In summary, the novel indel mutation within the goat PITX2 gene was significantly associated with litter size and growth traits, which could be used as an effective molecular marker for improving economic traits in the goat industry. The molecular mechanism is limited in this study, and it will be verified by subsequent studies.
The original data of the paper are available upon request from the corresponding author.
HY, EJ, and LQ designed the study and wrote the paper. HY and EJ performed the experiments. HY did the statistical analysis. HZ, LH and JL collected the samples. LQ edited and reviewed the paper.
The authors declare that they have no conflict of interest.
This work was funded by the National Natural Science Foundation of China
(nos. 31760650 and 31702115), Provincial Key Projects of Shaanxi (no. 2014KTDZ02-01),
Applied Basic Research Program of Qinghai Province (2014-ZJ-710), and West
Light Foundation of The Chinese Academy of Sciences. We greatly thank the
staff of the Shaanbei white cashmere goat breeding farm, Shaanxi province, P.R.
China for their collecting samples.
Edited by: Steffen Maak
Reviewed by: two anonymous referees
Aljanabi, S. M. and Martinez, I.: Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques, Nucleic Acids Res., 25, 4692–4693, 1997.
Angela, T., Mohsin, A. F. K., Nidia, M. M. O., Torlopp, A., Khan, M. A., Oliveira, N. M., Lekk, I., Soto-Jiménez, L. M., Sosinsky, A., and Stern, C. D.: The transcription factor Pitx2 positions the embryonic axis and regulates twinning, eLife, 3, e03743, https://doi.org/10.7554/eLife.03743, 2014.
Basu, M. and Roy, S. S.: Wnt∕β-Catenin Pathway is regulated by PITX2 homeodomain protein and thus contributes to the proliferation of human ovarian adenocarcinoma cell SKOV-3, J. Biol. Chem., 288, 4355–4367, https://doi.org/10.1074/jbc.M112.409102, 2013.
Bertko, E., Klammt, J., Dusatkova, P., Bahceci, M., Gonc, N., Ten, H. L., Kandemir, N., Mansmann, G., Obermannova, B., Oostdijk, W., Pfäffle, H., Rockstroh, L. D., Schlicke, M., Tuzcu, A. K., and Pfäffle, R.: Combined pituitary hormone deficiency due to gross deletions in the POU1F1 (PIT-1) and PROP1 genes, J. Hum. Genet., 62, 755–762, 2017.
Chen, L. and Gage, P. J.: Heterozygous Pitx2 null mice accurately recapitulate the ocular features of Axenfeld-Rieger syndrome and congenital glaucoma, Invest. Ophth. Vis. Sci., 57, 5023–5030, https://doi.org/10.1167/iovs.16-19700, 2016.
Chen, R., Yu, S., Ren, F., Lv, X. Y., and Pan, C. Y.: Detection of one large insertion/deletion (indel) and two novel SNPs within the SPEF2 gene and their associations with male piglet reproduction traits, Arch. Anim. Breed., 59, 275–283, https://doi.org/10.5194/aab-59-275-2016, 2016.
Cieslak, J., Flisikowska, T., Schnieke, A., Kind, A., Szydlowski, M., Switonski, M., and Flisikowski, K.: Polymorphisms in the promoter region of the adiponectin (ADIPOQ) gene are presumably associated with transcription level and carcass traits in pigs, Anim. Genet., 44, 340–343, https://doi.org/10.1111/j.1365-2052.2012.02397.x, 2013.
Cui, Y., Yan, H., Wang, K., Xu, H., Zhang, X., Zhu, H., Liu, J., Qu, L., Lan, X., and Pan, C.: Insertion/Deletion within the KDM6A gene is significantly associated with litter size in goat, Front. Genet., 9, 91, https://doi.org/10.3389/fgene.2018.00091, 2018.
Cui, Y., Zhang, Y., Wei, Z., Gao, J., Yu, T., Chen, R., Lv, X., and Pan, C.: Pig KDM5B: mRNA expression profiles of different tissues and testicular cells and association analyses with testicular morphology traits, Gene, 650, 27–33, https://doi.org/10.1016/j.gene.2018.01.092, 2018b.
Diana, E., Hsiao, Y. M., Michael, K. G., and Chrissa, K.: Gene networks during skeletal myogenesis, Isrn Developmental Biology, 2013, 348704, https://doi.org/10.1155/2013/348704, 2013.
El-Bayomi, K. M., Saleh, A. A., Awad, A., El-Tarabany, M. S., El-Qaliouby, H. S., Afifi, M., El-Komy, S., Essawi, W. M., Almadaly, E. A., and El-Magd, M. A.: Association of CYP19A1 gene polymorphisms with anoestrus in water buffaloes, Reprod Fertil Dev., 30, 487–497, https://doi.org/10.1071/RD16528, 2018.
Hassed, S. J., Li, S., Xu, W., and Taylor, A. C.: A novel mutation in PITX2 in a patient with Axenfeld-Rieger syndrome, Molecular Syndromology, 8, 7–9, https://doi.org/10.1159/000454963, 2017.
Hernandez, T. F., Rodríguez, O. L., Franco, D., and Aranega, A. E.: Pitx2 in embryonic and adult myogenesis, Frontiers in Cell & Developmental Biology, 5, 46, https://doi.org/10.3389/fcell.2017.00046, 2017.
Jin, Y., Cai, H., Liu, J., Lin, F., Qi, X., Bai, Y., Lei, C., Chen, H., and Lan, X.: The 10 bp duplication insertion/deletion in the promoter region within paired box 7 gene is associated with growth traits in cattle, Arch. Anim. Breed., 59, 469–476, https://doi.org/10.5194/aab-59-469-2016, 2016.
Lan, X. Y., Pan, C. Y., Chen, H., Zhang, C. L., Li, J. Y., Zhao, M., Lei, C. Z., Zhang, A. L., and Zhang, L.: An AluI PCR-RFLP detecting a silent allele at the goat POU1F1 locus and its association with production traits, Small. Ruminant. Res., 73, 8–12, https://doi.org/10.1016/j.smallrumres.2006.10.009, 2007.
L'Honoré, A., Ouimette, J. F., Lavertu, J. M., and Drouin, J.: Pitx2 defines alternate pathways acting through MyoD during limb and somitic myogenesis, Development, 137, 3847–3856, https://doi.org/10.1242/dev.053421, 2010.
Li, J., Erdenee, S., Zhang, S. L., Wei, Z. Y., Zhang, M., Jin, Y. Y., Wu, H., Chen, H., Sun, X. Z., Xu, H. W, Cai, Y., and Lan, X. Y.: Genetic effects of PRNP gene insertion/deletion (indel) on phenotypic traits in sheep, Prion, 12, 42–53, https://doi.org/10.1080/19336896.2017.1405886, 2018.
Li, N., Dobrev, D., and Wehrens, X. H.: PITX2: a master regulator of cardiac channelopathy in atrial fibrillation, Cardiovasc. Res., 109, 345–347, https://doi.org/10.1093/cvr/cvw008, 2016.
Margaret, B. and Peter, W. J. R.: Gene regulatory networks and transcriptional mechanisms that control myogenesis, Dev. Cell., 28, 225–238, https://doi.org/10.1016/j.devcel.2013.12.020, 2014.
Maryam, M., Javad, A., Rana, J. E., and Bahman, Y.: The roles of Wnt∕β-catenin pathway in tissue development and regenerative medicine, J. Cell. Physiol., 233, 5598–5612, https://doi.org/10.1002/jcp.26265, 2018.
Pak, S., Joel, S., Bader, I. C., O'Donovan, M., and Owen, M.: DNA poling: a tool for largeccal association studies, Nature Publishing Group, 3, 862–871, 2002.
Qiu, H. B., Sun, W. D., Yang, X., Jiang, Q. Y., Chen, S., and Lin, D. G.: Promoter mutation and reduced expression of BRCA1 in canine mammary tumor, Res. Vet. Sci., 103, 143–148, https://doi.org/10.1016/j.rvsc.2015.10.003, 2015.
Ren, F., Yu, S., Chen, R., Lv, X. Y., and Pan, C. Y.: Identification of a novel 12-bp insertion/deletion (indel) of iPS-related Oct4 gene and its association with reproductive traits in male piglets, Anim. Reprod. Sci., 178, 55–60, https://doi.org/10.1016/j.anireprosci.2017.01.009, 2017.
Ren, G., Chen, H., Zhang, L. Z., Lan, X. Y., Wei, T. B., Li, M. J., Jing, Y. J., Lei, C. Z., and Wang, J. Q.: A coding SNP of LHX4 gene is associated with body weight and body length in bovine, Mol. Biol. Reo., 37, 417–422, https://doi.org/10.1007/s11033-009-9486-6, 2010.
Sambrook, J. and Russell, D. W.: Molecular cloning: a laboratory manual, 3rd edition, Vol. 3, Coldspring Harbour Laboratory Press, UK, 2001.
Syeda, F., Kirchhof, P., and Fabritz, L.: PITX2-dependent gene regulation in atrial fibrillation and rhythm control, J. Physiol., 595, 4019–4026, https://doi.org/10.1113/JP273123, 2017.
Tessa, G. M. and Alexander, F. S.: Vg1-Nodal heterodimers are the endogenous inducers of mesendoderm, eLife, 6, e28183, https://doi.org/10.7554/eLife.28183, 2017.
Wang, K., Yan, H. L., Xu, H., Yang, Q., Zhang, S. H., Pan, C. Y., Chen, H., Zhu, H. J., Liu, J. W., Qu, L., and Lan, X. Y.: A novel indel within goat CSN1S1 gene significantly affects litter size, Gene, 671, 161–169, https://doi.org/10.1016/j.gene.2018.05.119, 2018.
Wang, X. Y., Yang, Q., Wang, K., Zhang, S. H., Pan, C. Y., Chen, H., Qu, L., Yan, H. L., and Lan, X. Y.: A novel 12-bp indel polymorphism within the GDF9 gene is significantly associated with litter size and growth traits in goats, Animal Genetics, 48, 735–736, https://doi.org/10.1111/age.12617, 2017.
Welsh, I. C., Thomsen, M., Gludish, D. W., Alfonso, P. C., Bai, Y., Martin, J. F., and Kurpios, N. A.: integration of left-right Pitx2 transcription and Wnt signaling drives asymmetric gut morphogenesis via Daam2, Dev. Cell, 26, 629, https://doi.org/10.1016/j.devcel.2013.07.019, 2013.
Yan, H., Zhang, F., Wang, K., Liu, J., Zhu, H., Pan, C., and Qu, L.: A novel 12 bp deletion within goat LHX4 gene significantly affected litter size, Arch. Anim. Breed., 61, 1–8, https://doi.org/10.5194/aab-61-1-2018, 2018.
Yang, Q., Yan, H. L., Li, J., Xu, H., Wang, K., Zhu, H. J., Chen, H., Qu, L., and Lan, X. Y.: A novel 14-bp duplicated deletion within goat GHR gene is significantly associated with growth traits and litter size, Anim. Genet., 48, 499–500, https://doi.org/10.1111/age.12551, 2017.
Yeonsoo, Y., Tingting, H., Giovane, G. T., Maki, W., Anna, K. H., Richard, R. B., Jaime, A., and Rivera, P.: Extra-embryonic Wnt3 regulates the establishment of the primitive streak in mice, Dev. Biol., 403, 80–88, https://doi.org/10.1016/j.ydbio.2015.04.008, 2015.
Yin, H. F., Fang, X. Y., Jin, C. F., Yin, J. F., Li, J. Y., Zhao, S. J., Miao, Q., and Song, F. W.: Identification of a novel frameshift mutation in PITX2 gene in a Chinese family with Axenfeld-Rieger syndrome, J. Zhejiang Univ-Sci B (Biomed & Biotechnol), 15, 43–50, https://doi.org/10.1631/jzus.B1300053, 2014.
Yusuke, M., Wnt∕β-catenin signaling and body plan formation in mouse embryos, Semin. Cell Dev. Biol., 17, 175–184, https://doi.org/10.1016/j.semcdb.2006.04.003, 2006.
Zhang, S. H., Xu, H., Liu, X. F., Yang, Q., Pan, C. Y., Lei, C. Z., Dang, R. H., Chen, H., and Lan, X. Y.: The muscle development transcriptome landscape of ovariectomized goat, Roy. Soc. Open Sci., 4, 1714–1715, https://doi.org/10.1098/rsos.171415, 2017.
Zhang, S. H., Xu, H., Kang, Z. H., Cai, H. F., Dang, R. H., Lei, C. Z., Chen, H., Guo, X., and Lan, X. Y.: Bovine PITX2 gene: mRNA expression analysis and association ananlysis, Gene, 669, 1–7, https://doi.org/10.1016/j.gene.2018.05.083, 2018.
Zhang, X. Y., Zhao, H. Y., Zhang, S. H., Zhang, S. H., Lei, C. Z., Chen, H., and Lan, X. Y.: Molecular cloning, sequence analysis and mRNA expression profiling of paired-liked homodomain transcription factor 2 gene (PITX2) in dairy goat (capra hircus), J. Agricult. Biotechnol., 23, 905–916, 2015.
Zhang, Y. H., Cui, Y., Zhang, X. L., Wang, Y. M., Gao, J. Y., Yu, T., Lv, X. Y., and Pan, C. Y.: Pig StAR: mRNA expression and alternative splicing in testis and Leydig cells, and association analyses with testicular morphology traits, Theriogenology, 118, 46–56, https://doi.org/10.1016/j.theriogenology.2018.05.031, 2018.
Zhao, H., He, S., Zhu, Y., Cao, X., Luo, R., Cai, Y., Xu, H., and Sun, X.: A novel 29 bp insertion/deletion (indel) variant of the LHX3 gene and its influence on growth traits in four sheep breeds of various fecundity, Arch. Anim. Breed., 60, 79–85, https://doi.org/10.5194/aab-60-79-2017, 2017.
Zhao, H. Y., Wu, X. F., Cai, H. F., Pan, C. Y., Lei, C. Z., Chen, H., and Lan, X. Y.: Genetic variants and effects on milk traits of the caprine paired-like homeodomain transcription factor 2 (PITX2) gene in dairy goats, Gene, 532, 203–210, https://doi.org/10.1016/j.gene.2013.09.062, 2013.
Zhou, F., Yang, Q., Lei, C. Z., Chen, H., and Lan, X. Y.: Relationship between genetic variants of POU1F1, PROP1, IGFBP3, genes and milk performance in Guanzhong dairy goats, Small Ruminant Res., 140, 40–45, https://doi.org/10.1016/j.smallrumres.2016.05.015, 2016.
Zhou, M., Liao, Y., and Tu, X.: The role of transcription factors in atrial fibrillation, J. Thorac. Dis., 7, 152, https://doi.org/10.1093/cvr/cvt261, 2015.