the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Effect of Mucin13 gene polymorphism on diarrhoeagenic E. coli adhesion pattern and its expression analysis in native Indian pigs
Rebeka Sinha
Nihar Ranjan Sahoo
Kush Shrivastava
Pushpendra Kumar
Salauddin Qureshi
Amit Kumar
Gandham Venkata Papa Pydi Siva Ravi Kumar
Bharat Bhushan
We identified genetic polymorphism in Mucin13 gene affecting E. coli adhesion patterns using (local isolate) diarrhoeagenic E. coli in Indian desi pigs. Five SNPs and one indel previously reported to be associated with enterotoxigenic E. coli (ETEC) F4ab/ac adhesion pattern were examined by designing PCR-RFLP protocol. The genotypic frequencies of only one SNP (g.22304A > G) differed significantly (at P≤0.05) in adhesive, non-adhesive and weakly adhesive population. The AA (306 sbp, 231 bp), AG (306, 231, 108, 198 bp) and GG (231, 198 bp, 108 b) genotypes of g.22304A > G locus were observed with frequencies 50.0 %, 21.25 % and 28.75 %, respectively and AG genotype was significantly (P≤0.05) associated with a non-adhesive pattern. The polymorphism information content of SNPs ranged from 17.67 (g.22124T > C) to 37.36 % (g.21471C > T) loci. Three loci (g.21471C > T, g.22124T > C and g.22304A > G) were significantly departed from Hardy–Weinberg equilibrium. The linkage disequilibrium analysis revealed locus g.22124T > C and g.22304A > G were significantly (P≤0.05) associated with each other. Expression profiling of target gene in jejuna of animals having AA, AG and GG genotypes revealed differences in various genotypes with the highest in the AA, moderate in the GG and low levels in the AG genotype, although they were statistically non-significant (at P≤0.05). The absence of significant effect of genotypes on MUC13 mRNA expression indicates no direct functional role, although the structural role can not be ignored as the putative receptor gene is located within targeted genomic region. Further, reports of same SNP association with an ETEC F4ab/ac adhesion pattern indicate the target gene's role in diarrhoea even caused by other strains of E. coli which is not ETEC.
- Article
(826 KB) - Full-text XML
- BibTeX
- EndNote
Piglet diarrhoea, which has consistently been identified as a primary infectious disease leading to poor health and mortality, accounts for 11.5 %–29.5 % of piglet deaths worldwide (Li et al., 2007). Among the potential bacterial causes, Escherichia coli are important pathogens in swine contributing about 56.2 % of the incidence and 24.7 % of mortality from diarrhoea (Shi, 2003). This microorganism adheres to receptors on intestinal brush borders of the host through its fimbriae and liberates enterotoxins that stimulate small intestine for secretion of massive fluid into gut lumen resulting in diarrhoea (Jin and Zhao, 2000; Van den Broeck et al., 2000). Pathotypes of E. coli that cause diarrhoea are referred to as diarrhoeagenic E. coli and are classified as per their clinical pathology, mechanism of disease induction and colonization site (Gomes Tania et al., 2016). Adhesion to intestinal epithelial cells, an essential prerequisite for the bacteria to cause diarrhoea is not equally found in all piglets as certain piglets are innately resistant (Sellwood et al., 1975). This differential adhesion pattern was found to be genetically controlled and inherited in dominant fashion in piglets (Bijlsma et al., 1982) and it may help in identifying the genetic cause for susceptibility.
While the exact/specific gene(s) that encodes for the resistance/susceptibility to enterotoxigenic E. coli (ETEC) are not yet known, the locus encoding the intestinal receptor was reported to be mapped to q41 region on pig chromosome 13 (Sus scrofa chromosome; 13) by several independent linkage analyses (Python et al., 2002; Jorgensen et al., 2003; Jacobsen et al., 2010). Among the putative positional candidate genes found in this region, studies in a collection of diverse outbred populations strongly supported Mucin13 (MUC13) to be the most likely responsible gene. A battery of 10 SNPs in MUC13 gene was reported to be significantly associated with the differential adhesion pattern with respect to ETEC F4ab/ac (Zhang et al., 2008; Ren et al., 2012). Furthermore, MUC13 gene was reported to encode two transcripts: MUC13A and MUC13B. Both have the characteristic PTS (proline, threonine and serine) regions enriched in distinct tandem repeats. MUC13B is predicted to be heavily O-glycosylated, forming the binding site of the bacterium while MUC13A does not have the O-glycosylation binding site (Ren et al., 2012), thus preventing adhesion. The presence of an insertion/deletion of 68 bp at intron 2 distinguishing MUC13A and MUC13B alleles was reported (Ren et al., 2012) and proposed to be used as a diagnostic marker. Neonatal diarrhoea is one of the most common causes of morbidity and mortality among piglets, causing heavy economic losses in the pork industry, especially in nations showing rapid agricultural development like India (Shome et al., 2005; Sinha et al., 2018), and E. coli is reported to be associated with severe diarrhoea, causing mortality in piglets early in life (Kumar and Soman, 2001; Mandakini et al., 2015). However, there is scarcity of literature about the genetic status of these receptors controlling the adhesion pattern in native pig population, which, if explored, can be utilized for enhancing genetic resistance against piglet diarrhoea. Previous studies have shown a differential jejunal expression of MUC13 gene in native Indian pigs and its potential important role in adhesion of E. coli (Sinha et al., 2018). The present investigation explored the genetic polymorphism in native Indian pigs with respect to indel/SNPs of MUC13, their association with an E. coli adhesion pattern (using local isolate) and effect of associated SNPs on its expression pattern.
2.1 Experimental animals and tissue collection
A total of 80 native pigs slaughtered in Bareilly, UP, India, were screened for genetic polymorphism in porcine MUC13 gene and E. coli adhesion pattern. Jejunum tissue samples were collected within 30 min post slaughter from pigs and brought to laboratory in ice under sterile conditions. After cleaning it properly with a cold phosphate buffer saline (PBS, pH 7.4), a small part (250 mg) of the sample was preserved in RNA later solution at −20 ∘C, muscle sample of 500 mg was used for DNA extraction and a small piece of jejunum (approximately 2 cm) was used for microscopic adhesion test (MAT) on the same day.
2.2 Diarrhoeagenic E. coli isolation, ETEC typing, biochemical and molecular characterization
The bacterial strain (E. coli) used for adhesion assay was isolated from the diarrhoeic piglets of a swine production farm of ICAR-Indian Veterinary Research Institute. They were characterized biochemically as well as by amplification and sequencing of partial 16S ribosomal RNA gene (gb|KJ810542), before use for an adhesion test. This isolate was cultured in a brain–heart infusion (BHI) agar plate overnight at 37 ∘C, and a single colony was incubated in lysogeny broth (LB) medium (Tryptone, Yeast extract, NaCl, pH 7.0–7.2) at 37 ∘C for 16–18 h at 180 rpm. Once the final OD reached 1.0 at 520 nm, the culture was kept at 4 ∘C for use on the same day.
2.3 Microscopic adhesion test (MAT)
Samples were tested for adhesion pattern through MAT following Li et al. (2007). Briefly, for preparation of the epithelial cells a piece of jejunum (approx. 2 cm) was excised from the small intestine, cut open along the longitudinal axis and washed with a cold PBS (pH 7.4) containing 0.1 M EDTA to make free of intestinal contents. Subsequently, the sample was immersed in a cold hypotonic EDTA solution (5 mM EDTA, adjusted to pH 7.4) for 20 min with gentle rinsing. Epithelial cells were removed from mucosal surface of jejunum and immersed in the cold hypotonic EDTA solution for 30 min. Then enterocytes were homogenized and filtered through a clean muslin cloth. The filtrate was centrifuged at 3500 rpm for 10 min to form pellets, and the cells were resuspended in 5–6 mL cold PBS. Brush border suspension was added with 100 µL gentamicin sulfate (1 mg mL−1) and sodium azide (3 mM) to make a final concentration of 1×106 cells mL−1 and stored at 40 ∘C for use on the same day. For the adhesion test, equal volume of brush border cell suspension and fresh bacterial suspension (100 µL each) were mixed in a tissue culture plate and incubated for 30 min at 37 ∘C at 200 rpm. A drop of the mixed suspension was examined for the adhesion pattern using light microscope under 40× objective. A single epithelial cell was considered adhesive when there were more than five bacterial cells adhering to the brush border membrane. Twenty well-separated and intact enterocytes were examined from each animal, and the animals were classified as adhesive, non-adhesive and weakly adhesive phenotypes (Li et al., 2007).
2.4 Design of PCR-RFLP protocol and DNA extraction
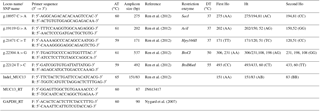
SNP loci were selected based on previous reports of association with an ETECF4 adhesion pattern (Ren et al., 2012), and oligonucleotide primers flanking SNPs in MUC13 gene (GenBank accession no. JN613418.1) were designed using the online software Primer3 plus tool (http://primer3plus.com/cgi-bin/dev/primer3plus.cgi, last access: 25 October 2013) (Table 1). Suitable restriction enzymes were selected for each SNP loci with the help of the NEB cutter V2.0 online software (http://www.tools.neb.com/NEBcutter2, last access: 25 October 2013). Genomic DNA was isolated from muscle tissue (500 mg) by phenol : chloroform : isoamyl alcohol (PCI) extraction method (Sambrook and Russell, 2012). Briefly, about 500 mg muscle tissue was minced with a homogenizer and added with a DNA extraction buffer (1 % 1 M Tris pH 8.0; 8 % 5 M NaCl 0.4 % 0.5 M EDTA) in a ratio of 3:1. The cell debris was digested by proteinase K (20 mg mL−1) added in two pulses along with 10 % SDS under overnight incubation at 65 ∘C. The following morning, the content was extracted with an equal amount of equilibrated phenol (Tris saturated phenol pH>7.8), () and chloroform:isoamyl alcohol (24:1). The precipitation of DNA was done by double volume of isopropanol (at room temperature) after addition of sodium acetate (3 M). The DNA pellet was washed twice in 70 % ethanol, air dried and re-suspended in TE (Tris-HCl 10 mM, EDTA 0.5 M, pH 8.0) buffer. The quality, integrity and concentration of the genomic DNA were checked by using horizontal submarine agarose gel (0.8 %) electrophoresis and NanoDrop spectrophotometer (ND 1000).
2.5 Mass genotyping using PCR-RFLP and indel typing
All animals were genotyped for SNPs and indel. The PCR amplification was carried out in a 25 µL reaction mix, containing 50 ng of porcine genomic DNA as a template, 1× PCR buffer, 1.0 mM of forward and reverse primers each, 200 mM of each dNTP, 2.5 mM MgCl2 and 1 U of Taq polymerase (Fermentas). The reactions were performed on the ABI thermal cycler (Veriti, Applied Biosystems, USA) under the thermal cycle profile; denaturation at 95 ∘C for 10 min in the first cycle followed by 35 cycles of 95 ∘C for 45 s, 59–61 ∘C (Table 1) for 45 s, 72 ∘C for 45 s; and final extension at 72 ∘C for 10 min for the last cycle. The amplified products were checked using 1.5 % agarose gel electrophoresis in 1× TBE buffer along with a 100 bp DNA ladder (GeneRuler, Thermo Scientific). Each amplified product was digested with 1 IU of different restriction enzyme using the manufacturer's recommended buffer and conditions for digestion (Table 1). The digested PCR products were subjected to electrophoresis in 2.5 % w∕v agarose gel (low EEO) for 1 h at 120 V against 100 and/or 50 bp DNA ladder and documented under a gel documentation system. Each individual revealed a particular pattern or RFLP genotype depending on presence and absence of RE sites. The indel located at intron 2 (Ren et al., 2012), distinguishing the MUC13A and MUC13B alleles, was genotyped by direct amplification to yield amplified products of 83 bp (MUC13B) and 151bp (MUC13A) with the reported primers (Table 1).
2.6 Expression profiling
Four animals from each different genotypes (AG, AA and GG) of the SNP significantly associated with an adhesive pattern (g22304A > G) were studied for the expression profiling of MUC13 gene in their jejuna. Total RNA was isolated from target tissue using TRIzol Reagent (Sambrook and Russell, 2012). The RNA was precipitated, washed and re-suspended in nuclease-free water. The first strand cDNA was synthesized from isolated good-quality () total RNA using RevertAid H Minus First Strand cDNA Synthesis Kit (Thermo Scientific) as per the manufacturer's instructions. The resulting complementary DNA (cDNA) was checked by PCR with porcine GAPDH (glyceraldehyde-3-phosphate dehydrogenase) primers (Nygard et al., 2007) and used in quantitative RT-PCR (qRT-PCR) reactions. The primer efficiencies (Table 1) of target and housekeeping genes were determined by running a standard curve for each assay by levels of six serial dilutions of cDNA containing the template. qRT-PCR was performed using SsoFast EvaGreen® qPCR kit (Bio-Rad) using Stratagene Mx3000P (Agilent Technologies, USA) thermal cycler operated by MxPro™ QPCR software. A no template control (real-time master mix and primers) was put for checking contamination in the reaction components other than cDNA. The master mix was prepared using 8.0 µL of nuclease-free water, 0.50 µL of forward and reverse primers each; 10 µL of EvaGreen mix (Bio-Rad) and 1 µL of cDNA was added to the master mix. Three-segmented qPCR amplification programme was used (one cycle of Hot start PCR at 95 ∘C for 15 s followed by 35 cycles of denaturation at 95 ∘C for 10 s and annealing/extension at 60 ∘C for 12 s). The dissociation curve analysis was performed for one cycle at 95 ∘C for 1 min, 65 ∘C for 30 s, 65–95 ∘C (2 ) and 95 ∘C for 30 s.
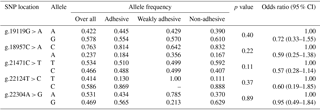
Table 2Genotype frequency distribution of porcine MUC13 gene polymorphism.
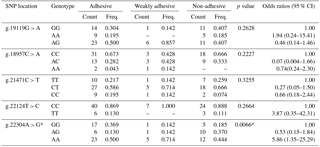
* Significant at P≤0.05; the positions of SNPs refer to GenBank accession no. JN613418.
2.7 Statistical analysis
The association between various allelic variants with an adhesion pattern was explored by using logistic model for categorical variables with the SAS 9.3 software (SAS® 9.3 Software Version 9.3.2011, 2011). The univariate analysis for logistic regression considered the adhesion status as a categorical response variable (adhesive, weakly adhesive and non-adhesive), and SNPs were included as possible explanatory variables. Data were analysed using PROC LOGISTIC procedure and odds ratios (ORs) with a 95 % confidence interval. The relative risk of incidence among the genotypes was analysed using a logistic regression model, , which compared heterozygous (Ht) and homozygous (Ho) genotypes and estimated one/two ORs, one for Ht and the other for Ho, where α, β1, β2 and γ are parameters. The PROC ALLELE procedure was used for testing of Hardy–Weinberg equilibrium (HWE), estimation of polymorphism information content (PIC) and heterozygosity of SNPs markers as well as to test linkage disequilibrium (LD) between any two SNP loci. For qRT-PCR, optical data were collected at end of each extension step and relative expression of PCR product was determined by the equation given by Pfaffl (2001). The statistical significance (P≤0.05) of differences in mRNA expressions of the examined factors was assessed by using one-way ANOVA.
3.1 Phenotyping of the animals
All the animals were screened for patterns of brush border adherence by E. coli using microscopic adhesion test (MAT). Out of 80 animals, 27 pigs were found to be negative for adhesion (i.e. non-adhesive) with E. coli, whereas 53 animals were positive for adhesion (i.e. adhesive). Among the positive samples 46 pigs were adhesive and 7 were weakly adhesive. The PCR-based fimbrial typing showed that the current isolate was ETEC F4ab/ac-negative.
3.2 Gene, genotype frequencies and their association with an adhesion pattern
The genotypic and allelic frequencies of target population (Tables 2 and 3) revealed all five loci were polymorphic with two (g.22124T > C) to three (g.19119G > A, g.18957C > A, g.21471C > T and g.22304A > G) genotypes. While four loci (g.19119G > A, g.18957C > A, g.21471C > T and g.22124T > C) showed non-significant association with an adhesion pattern, only one locus (g.22304A > G) showed a statistically significant (P≤0.05) association with an E. coli adhesion pattern. This locus had three genotypes, viz., GG, AG and AA, with their overall frequencies 0.288, 0.213 and 0.500, respectively. The PROC LOGISTIC procedure revealed all three genotypes (GG, AG and AA) of locus g.22304A > G had significantly different ORs between E. coli adhesive, non-adhesive and weakly adhesive pattern (Tables 2 and 3). The animals were also typed for an indel variation at intron 2 in MUC13 gene (Ren et al., 2012). However, our current population was found to be homozygous (MUC13B) for this variation, and hence further analysis was not carried out.
3.3 Hardy–Weinberg equilibrium (HWE) and linkage disequilibrium test
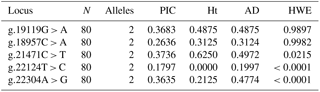
The PIC ranged from 0.179 (g.22124T > C) to 0.373 (g.21471C > T) and heterozygosity value ranged from 0.0 (g.22124T > C) to 0.6250 (g.21471C > T) in the population (Table 4). Test for HWE showed that three loci (g.21471C > T, g.22124T > C and g.22304A > G) significantly deviated from HWE, whereas others (g.22124T > C and g.22304A > G) were found to be in HWE. The linkage disequilibrium test using χ2 probabilities revealed locus g.22124T > C and g.22304A > G were significantly (at P≤0.05) associated with one another. None of the other loci showed significant LD with one another.
3.4 Expression profile of MUC13 across various genotypes
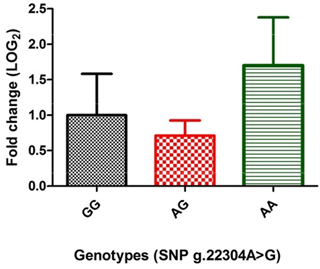
The expression analysis, to study the direct effect of genotypes of the significantly associated SNP (g.22304A > G), revealed that porcine MUC13 expression was different in different genotypes (Fig. 1). The level was highest in AA (1.699±0.67 fold) genotype, moderate in GG and low in the AG (0.712±0.21 fold) genotype and was found to be statistically non-significant (at P≤0.05) in both AG and AA genotype.
One of the important bacterial causes of piglet diarrhoea is E. coli infection. The colonization of E. coli to intestinal lumen causes release of enterotoxins, ultimately resulting in severe diarrhoea (Van den Broeck et al., 2000). The adhesion of E. coli affects colonization, which interferes with the disease process. The difference in pattern of adhesion and the genetic basis of this adhesion causing inherent resistance to the infection was targeted to be explored. Five SNP loci were selected based on previous reports by Ren et al. (2012) which were significantly associated with an ETECF4 adhesion pattern in a different outbred population. Only one SNP (g.22304A > G) was found to be significantly associated with an adhesion pattern of E. coli with intestinal epithelial cells. However, the pattern of adhesion was studied with respect to the locally isolated diarrhoeagenic E. coli, which were non-ETEC. The SNP on MUC13 gene had been previously mapped to SSC13q41 genomic region and also had been reported to be a candidate gene for E. coli resistance/susceptibility in piglets (Grange et al., 1998; Ren et al., 2012). Previous reports on MUC13 SNPs showed significant association for two SNPs (c.576C > T and c.935A > C) with E. coli resistance/susceptibility (Zhang et al., 2008) with respect to ETECF4. Out of three genotypes (GG, AG and AA) observed in this locus, genotype AA had significantly higher ORs of being associated with adhesiveness. However, the relative risk of incidence using a logistic regression model showed a non-significant association of either allele “G” or “A”, at locus g.22304A > G, with incidence or the adhesion pattern (Table 4). Our investigation showed genotype AG at locus g.22304A > G was associated with non-adhesive pattern of E. coli. Three out of five loci were significantly deviated from HWE. The significant departure from HWE may be ascribed due to natural selection or pre-born mortality in the population. The present investigation revealed that the locus which significantly affected the incidences of E. coli adhesion did not have significant LD with other locus except g.22124T > C, indicating its (g.22124T > C) usefulness for indirect selection. It infers that the chances of getting haplotype(s) from the loci significantly affecting E. coli adhesion pattern were rare.
An indel of 68 bp in intron 2 has been identified in MUC13 gene with the longer sequence for MUC13A and the shortened one for MUC13B and has been used as a diagnostic marker for typing these alleles (Ren et al., 2012). We used the same marker for genotyping our native pig population. However, all the animals were found to be carrying only MUC13B allele and hence were monomorphic for this marker.
Further to decipher whether there is any transcriptional level change due to SNPg.22304A > G in MUC13 gene expression, qRT-PCR was performed among various genotypes using MUC13 as a target gene and GAPDH as housekeeping control. Our previous results showed that the expression of MUC13 in jejuna tissue in these animals was different in adhesive, weakly adhesive and non-adhesive phenotypes. However, statistically this difference was non-significant (at P≤0.05) (Sinha et al., 2018). RT-PCR analysis revealed that although the porcine MUC13 mRNA expression was different in different genotypes (Fig. 1), with the highest level in genotype AA, moderate levels in the GG and low levels in the AG, the within-genotype difference was found to be statistically non-significant (at P≤0.05). However, previous studies on MUC13 expression by Schroyen et al. (2012) reported a significant difference in expression (at P≤0.05) between F4ac receptor-positive and F4ac receptor-negative animals.
The SNP g.22304A > G showed a statistically significant (at P≤0.05) association with an E. coli adhesion pattern using local diarrhoeagenic E. coli. The genotype AG at locus g.22304A > G was associated with a non-adhesive pattern. However, animals with AA, AG and GG genotypes revealed differences in expression level of MUC13, although they were statistically non-significant (at P≤0.05). Therefore, the investigation underscores the fact that porcine MUC13 gene may not be directly affecting resistance/susceptibility to diarrhoea due to E. coli in pigs. However, it certainly could form a positional genetic maker owing to its mapping location.
The research data can be accessed from ICAR-Indian Veterinary Research Institute, Animal Genetics Division, with the necessary formalities. As the institute is publicly funded, the data are publicly accessible in the host country. The data for the current research are part of the Institutional Project of Division of Animal Genetics, ICAR-IVRI, and can be made available on request to the corresponding author.
NRS designed the research programme and got the grant. RS, NRS, KS, PK, SQ and BB conducted the experiment. AK and GRK analysed data and interpreted the results. NRS and RS prepared the manuscript for communication and have contributed equally to the manuscript preparation.
The authors declare that they have no conflict of interest.
All necessary permissions from Institutional Animal Ethics Committee (IAEC) were obtained before experimentation.
The authors are thankful to the Indian Council of Agricultural Research (ICAR)
for providing necessary financial assistance in the form of an institute
project (grant no. IXX10577) to the corresponding author, as well as a fellowship
to the first author during the study. They are also thankful to R. K. Singh, Director,
ICAR-IVRI, Izatnagar, Bareilly, India, for providing
necessary facilities to carry out this research work. Further, they are
thankful to owners of different slaughter houses in and around Bareilly for
facilitating collection of samples.
Edited by:
Steffen Maak
Reviewed by: M. S. Thakur and one anonymous
referee
Bijlsma, I. G., De Nijs, A., Van Der Meer, C., and Frik, J. F.: Different pig phenotypes affect adherence of Escherichia coli to jejunal brush borders by K88ab, K88ac, or K88ad antigen, Infect. Immun., 37, 891–894, 1982.
Gomes Tania, A. T., Elias Waldir, P., Scaletsky Isabel, C. A., Guth Beatriz, E. C., Rodrigues Juliana, F., Piazza Roxane, M. F., Ferreira Luis, C. S., and Martinez, M. B.: Diarrheagenic Escherichia coli, Braz. J. Microbiol., 47S, 3–30, 2016.
Grange, P. A., Erickson, A. K., Anderson, T. J., and Francis, D. H.: Characterization of the carbohydrate moiety of intestinal mucin-type sialoglycoprotein receptors for the K88ac fimbrial adhesin of Escherichia coli, Infect. Immun., 66, 1613–1621, 1998.
Jacobsen, M., Kracht, S. S., Esteso, G., Cirera, S., Edfors, I., Archibald, A., Bendixen, C., Andersson, L., Fredholm, M., and Jorgensen, C. B.: Refined candidate region specified by haplotype sharing for Escherichia coli F4ab/F4ac susceptibility alleles in pigs, Anim Genet., 41, 21–25, 2010.
Jin, L. Z. and Zhao, X.: Intestinal receptors for adhesive fimbriae of enterotoxigenic Escherichia coli (ETEC) K88 in swine – a review, Appl. Microbiol. Biotechnol., 54, 311–318, 2000.
Jorgensen, C. B., Cirera, S., Anderson, S. I., Archibald, A. L., Raudsepp, T., Chowdhary, B., Edfors-Lilja, I., Andersson, L., and Fredholm, M.: Linkage and comparative mapping of the locus controlling susceptibility towards E. coli F4ab/ac diarrhoea in pigs, Cytogenet Genome Res., 102, 157–162, 2003.
Kumar, R. and Soman, J. P.: Studies on Escherichia coli isolates from piglet diarrhoea, Indian Vet J., 78, 879–882, 2001.
Li, Y., Qiu, X., Li, H., and Zhang, Q.: Adhesive Patterns of Escherichia coli F4 in Piglets of Three Breeds, J. Genet. Genomics., 34, 591–599, 2007.
Mandakini, R., Dutta, T. K., Chingtham, S., Roychoudhury, P., Samanta, I., Joardar, S. N., Pachauau, A. R., and Chandra, R.: ESBL-producing Shiga-toxigenic E. coli (STEC) associated with piglet diarrhoea in India, Trop. Anim. Health Prod., 47, 377–381, 2015.
Nygard, A. B., Jorgenson, C. B., Cierra, S., and Fredholm, M.: Selection of reference genes for gene expression studies in pig tissues using SYBR green qPCR, BMC Mol. Biol., 8, 67, https://doi.org/10.1186/1471-2199-8-6, 2007.
Pfaffl, M. W.: A new mathematical model for relative quantification in real-time RT-PCR, Nucleic Acids Res., 29, 2002–2007, 2001.
Python, P., Jorg, H., Neuenschwander, S., Hagger, C., Stricker, C., Burgi, E., Bertschinger, H. U., Stranzinger, G., and Vogeli, P.: Fine-mapping of the intestinal receptor locus for enterotoxigenic Escherichia coli F4ac on porcine chromosome 13, Anim Genet., 33, 441–447, 2002.
Ren, J., Yan, X., Ai, H., Zhang, Z., Huang, X., Ouyang, J., Yang, M., Yang, H., Han, P., and Huang, L.: Susceptibility towards enterotoxigenic Escherichia coli F4ac diarrhoea is governed by the MUC13 gene in Pigs, PLoS One, 7, e44573, https://doi.org/10.1371/journal.pone.0044573, 2012.
Sambrook, J. and Russell, D. W.: Molecular Cloning, a Laboratory Manual (4th edn.), Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 2012.
SAS® 9.3 Software [computer program]: Version 9.3.2011, SAS Institute Inc, Cary, NC, SAS Institute Inc., 2011.
Schroyen, M., Stinckens, A., Verhelst, R., Cox, E., Niewold, T., and Buys, N.: Susceptibility of piglets to enterotoxigenic E. coli is not related to the expression of MUC13 and MUC20, Anim. Genet., 43, 324–327, 2012.
Sellwood, R., Gibbons, R. A., Jones, G. W., and Rutter, J. M.: Adhesion of enteropathogenic Escherichia coli to pig intestinal brush borders: the existence of two pig phenotypes, J. Med. Microbiol., 8, 405–411, 1975.
Shi, Q. S.: The review of the receptors of ETEC F4, Pigs and Poultry, 23, 33–35, 2003.
Shome, R., Shome, B .R., Rahman, M., Kumar, A., Murugkar, H. V., Rahman, H., and Bujarbaruah, K.: Plasmid diversity in Escherichia coli strains isolated from piglet diarrhoea, Indian J. Anim. Sci., 75, 196–198, 2005.
Sinha, R., Sahoo, N. R., Kumar, P., Qureshi, S., Kumar, A., Ravikumar, G. V. P. P. S., and Bhushan, B.: Comparative jejunal expression of MUC 13 in Indian native pigs differentially adhesive to diarrhoeagenic E. Coli, J. Appl. Anim. Res., 46, 107–111, 2018.
Van Den Broeck, W., Cox, E., Oudega, B., and Goddeeris, B. M.: The F4 fimbrial antigen of Escherichia coli and its receptors, Vet. Microbiol., 71, 223–244, 2000.
Zhang, B., Ren, J., Yan, X., Huang, X., Ji, H., Peng, Q., Zhang, Z., and Huang, L.: Investigation of the porcine MUC13 gene: isolation, expression, polymorphisms and strong association with susceptibility to enterotoxigenic Escherichia coli F4ab/ac, Anim. Genet., 39, 258–266, 2008.